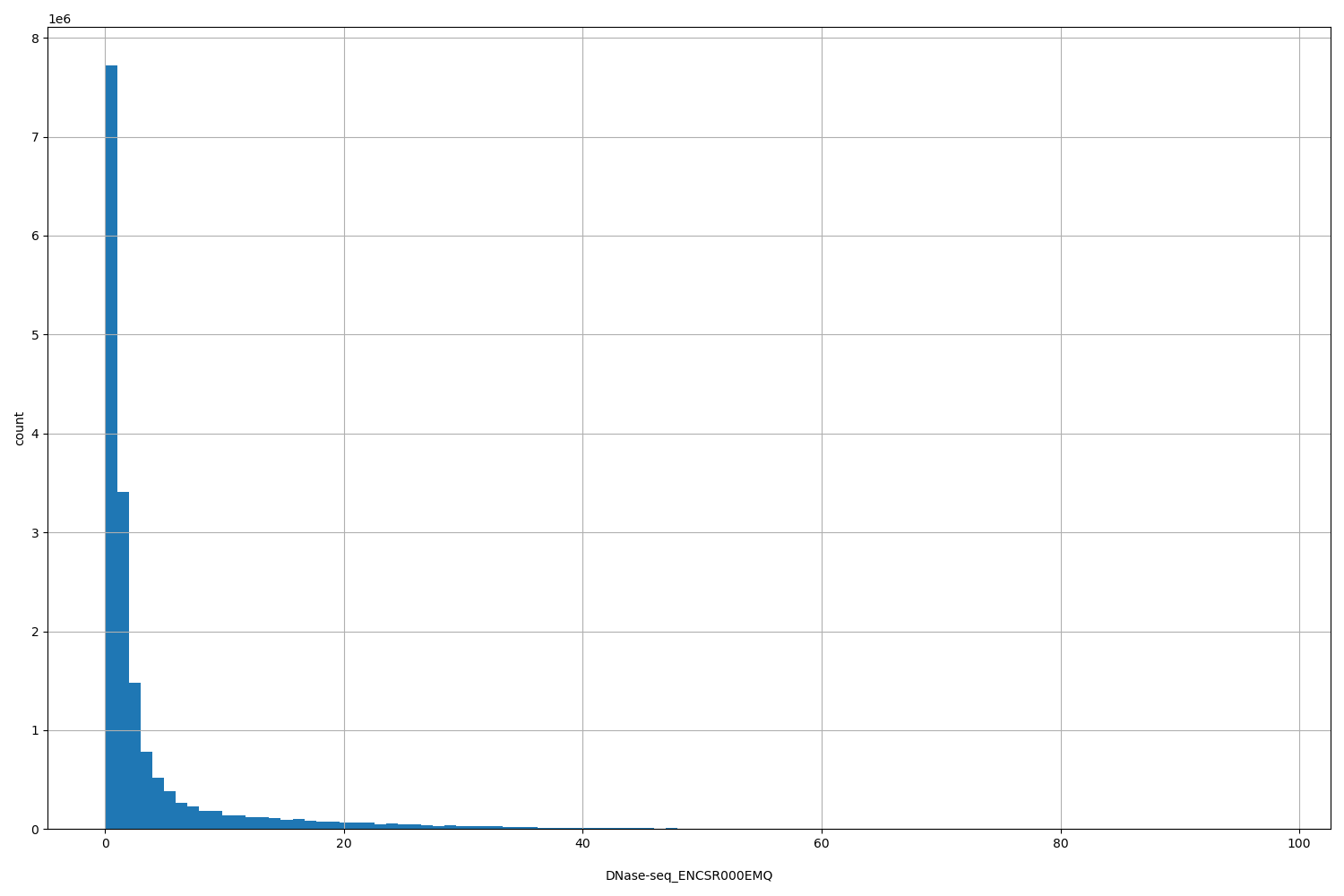
DNase-seq_ENCSR000EMQ
| Id: | DNase-seq/ENCSR000EMQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EMQ [biosample_summary="Homo sapiens GM06990"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_error: Alignment file {ENCFF311YYM|/files/ENCFF311YYM/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 16232001 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_internal_action: Released analysis {ENCAN517CSJ|/analyses/ENCAN517CSJ/} has in progress subobject document {59a15220-950a-456d-9cd3-77e9574cc42c|/documents/59a15220-950a-456d-9cd3-77e9574cc42c/} |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EMQ | float |
DNase-seq_ENCSR000EMQ |
DNase-seq ENCSR000EMQ [biosample_summary="Homo sapiens GM06990"]
|
 |
[0.0665, 97.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF239QAE.bed.gz | 894.8 KB | 58794c5bfe9aa9555aaa4d2bcfa871d7 |
| ENCFF239QAE.bed.gz.dvc | 100.0 B | cb4ab20e20c2adb9a83a7d599282b8f1 |
| ENCFF239QAE.tabix.bed.gz | 823.13 KB | a68e912c6f95dba818c638c80facb02f |
| ENCFF239QAE.tabix.bed.gz.dvc | 106.0 B | 21296569a16ec71b5cfb94b7c45029ab |
| ENCFF239QAE.tabix.bed.gz.tbi | 332.16 KB | 6534f677b6871d3af6c1778b62076e6f |
| ENCFF239QAE.tabix.bed.gz.tbi.dvc | 110.0 B | 34dc36ae1f279ea8a48294d915985365 |
| genomic_resource.yaml | 2.06 KB | e53ac60702426c2ca3a1aa6550a1f4c1 |
| genomic_resource_original.yaml | 1.99 KB | 58d52b507adc6e62af141841d550e902 |
| statistics/ |