DNase-seq_ENCSR000EMP
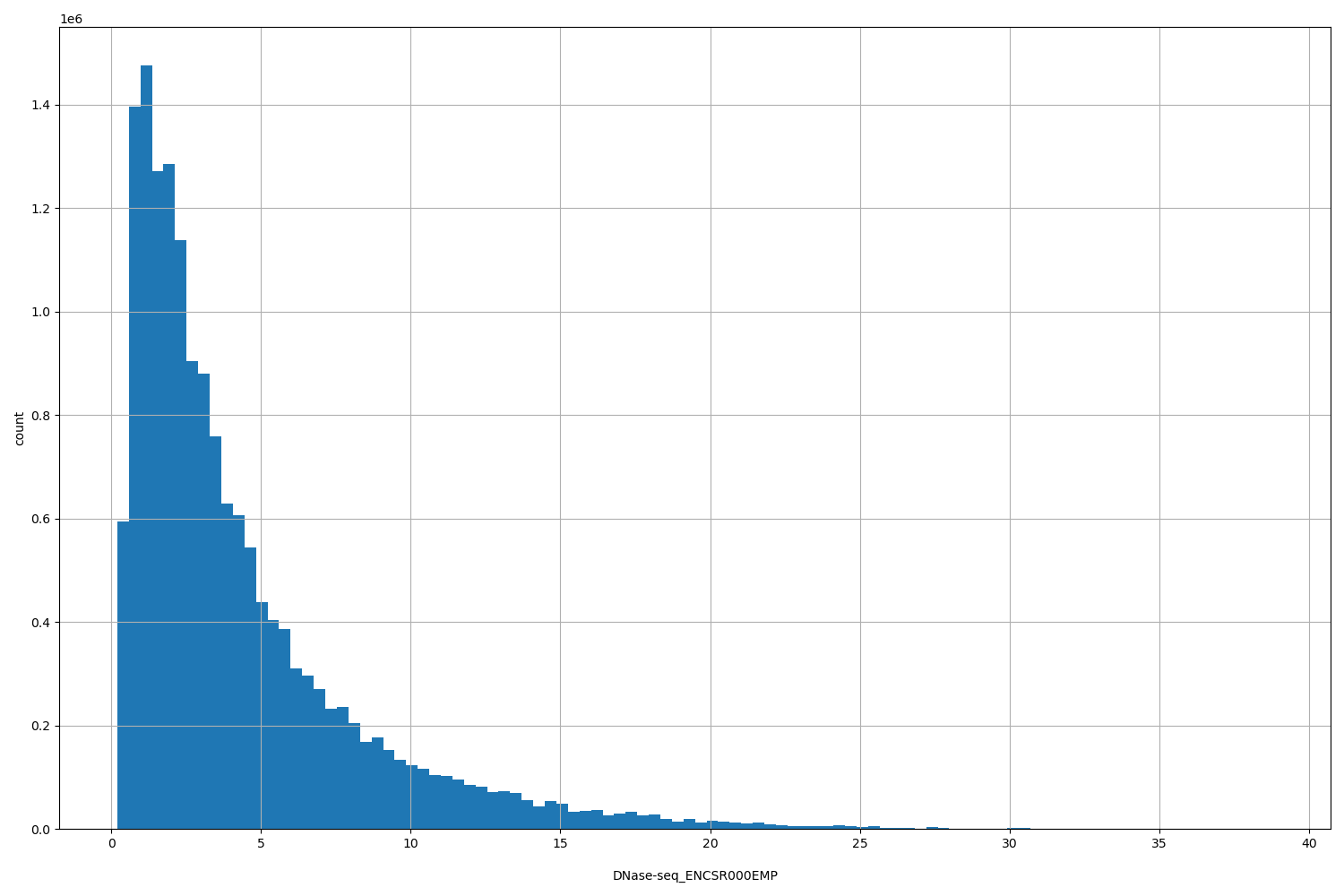
| Id: | DNase-seq/ENCSR000EMP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EMP [biosample_summary="Homo sapiens GM04504"] |
| Description: |
status: released biological_replicates: Rep 2 summary: output_type: peaks audit_warning: Alignment file {ENCFF708GUH|/files/ENCFF708GUH/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 30870575 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Alignment file {ENCFF928WMM|/files/ENCFF928WMM/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 30349249 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EMP | float |
DNase-seq_ENCSR000EMP |
DNase-seq ENCSR000EMP [biosample_summary="Homo sapiens GM04504"]
|
 |
[0.198, 38.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF041KUX.bed.gz | 968.72 KB | 727c27ab2d462ce7aa8045c57d0a5ffb |
| ENCFF041KUX.bed.gz.dvc | 100.0 B | fd9252005ca686a62f47888224560d65 |
| ENCFF041KUX.tabix.bed.gz | 863.13 KB | 63301ddf891fed6249da17df6dd020d2 |
| ENCFF041KUX.tabix.bed.gz.dvc | 106.0 B | d6d7e9da292b19201906170cdfd13191 |
| ENCFF041KUX.tabix.bed.gz.tbi | 434.53 KB | 0c60cb92c03c442f5b3ac4e15cdab644 |
| ENCFF041KUX.tabix.bed.gz.tbi.dvc | 110.0 B | b740b01562205f13db75a3ad39448f3c |
| genomic_resource.yaml | 2.35 KB | 413e19b6db21c147a83a933dbf65b52b |
| genomic_resource_original.yaml | 2.28 KB | cb1e9cd906c8c6202bb9737686efc5a9 |
| statistics/ |