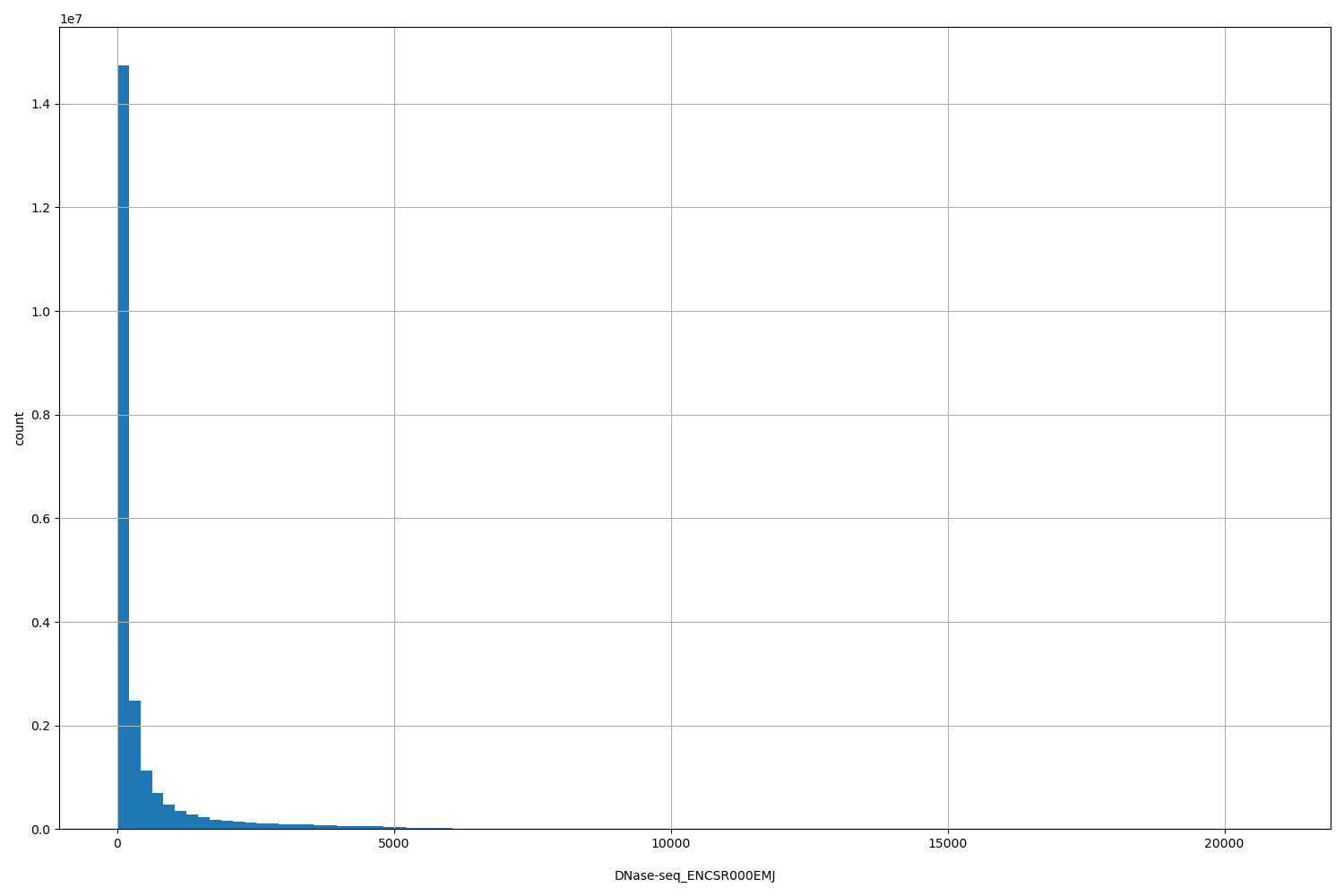
DNase-seq_ENCSR000EMJ
| Id: | DNase-seq/ENCSR000EMJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EMJ [biosample_summary="Homo sapiens B cell female adult (43 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (43 years) output_type: peaks audit_internal_action: Archived analysis {ENCAN556ZAU|/analyses/ENCAN556ZAU/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_not_compliant: Replicate concordance in DNase-seq experiments is measured by calculating the Pearson correlation between signal quantification of the replicates. ENCODE processed signal files {ENCFF801IWM|/files/ENCFF801IWM/}, {ENCFF305QCJ|/files/ENCFF305QCJ/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL001DNS|/pipelines/ENCPL001DNS/}) for GRCh38 assembly have a Pearson correlation of 0.89. According to ENCODE standards, in an isogenic assay a Pearson correlation value > 0.9 is recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/} ) audit_warning: Alignment file {ENCFF210XND|/files/ENCFF210XND/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 33733574 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EMJ | float |
DNase-seq_ENCSR000EMJ |
DNase-seq ENCSR000EMJ [biosample_summary="Homo sapiens B cell female adult (43 years)"]
|
 |
[3, 2.09e+04] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF541VKE.bed.gz | 1.42 MB | 00562290cbbe755d86f7701a4cd099c9 |
| ENCFF541VKE.bed.gz.dvc | 101.0 B | f1e792022d59a1ae223053ebd9ed8382 |
| ENCFF541VKE.tabix.bed.gz | 1.22 MB | 0ff7a183c21a0707f3f147fc3c38b549 |
| ENCFF541VKE.tabix.bed.gz.dvc | 107.0 B | 68a080fe8382ccd3424547143c0510f1 |
| ENCFF541VKE.tabix.bed.gz.tbi | 482.26 KB | 0eef197e018b31ed03753c411a47f4b6 |
| ENCFF541VKE.tabix.bed.gz.tbi.dvc | 110.0 B | 3c258962ff666db265ee157c6426bce4 |
| genomic_resource.yaml | 2.56 KB | 8734d1308bcc267994be8a7362d8fb8e |
| genomic_resource_original.yaml | 2.47 KB | be831b2fb17572da7aebed3ab54c8e88 |
| statistics/ |