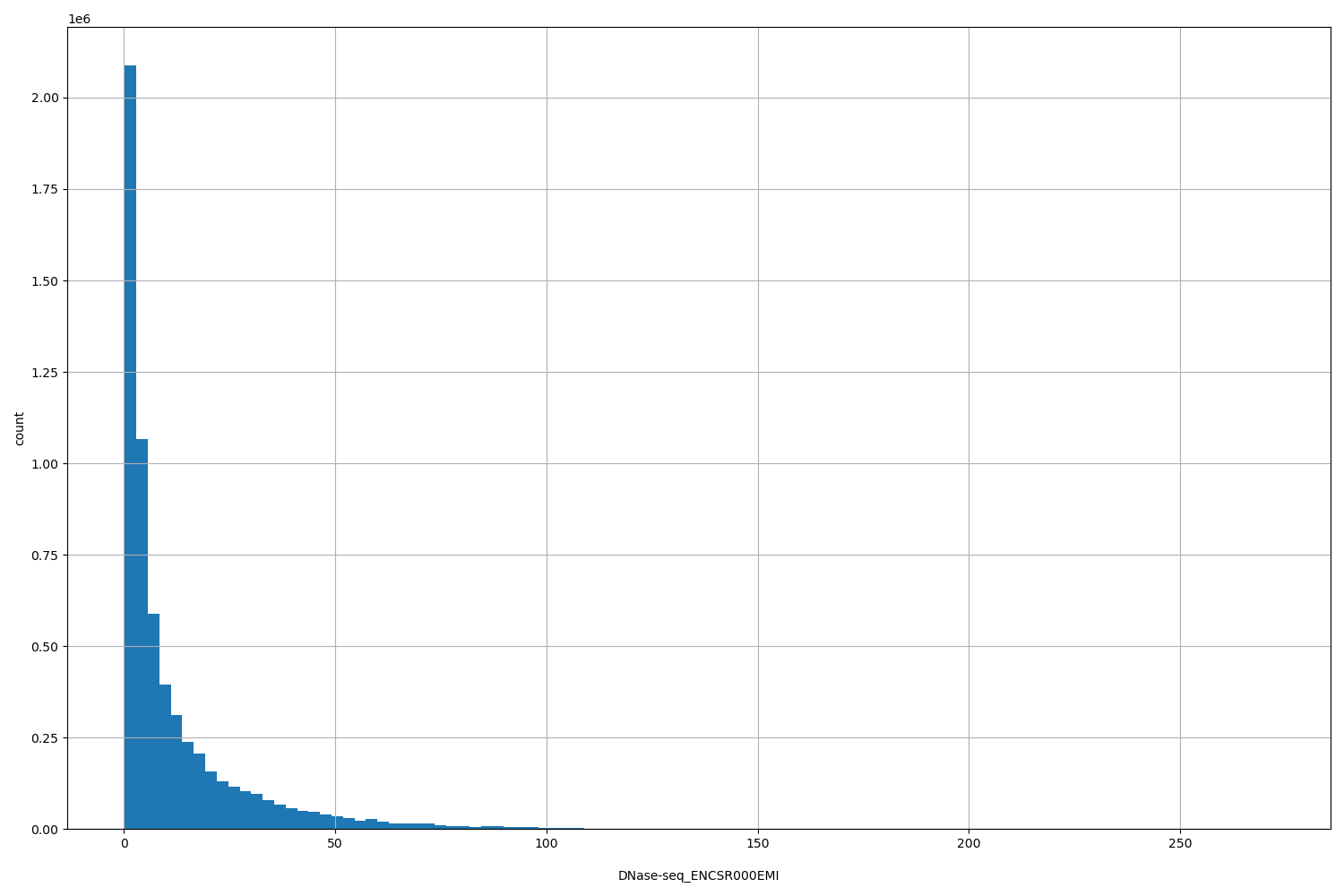
DNase-seq_ENCSR000EMI
| Id: | DNase-seq/ENCSR000EMI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EMI [biosample_summary="Homo sapiens Caco-2"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_internal_action: Released analysis {ENCAN306CUQ|/analyses/ENCAN306CUQ/} has in progress subobject document {29ecea40-1acd-4481-8342-6170e44cd7d1|/documents/29ecea40-1acd-4481-8342-6170e44cd7d1/} audit_warning: Alignment file {ENCFF895GQL|/files/ENCFF895GQL/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 22734571 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Alignment file {ENCFF474LBE|/files/ENCFF474LBE/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 20493023 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EMI | float |
DNase-seq_ENCSR000EMI |
DNase-seq ENCSR000EMI [biosample_summary="Homo sapiens Caco-2"]
|
 |
[0.265, 272] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF175ACH.bed.gz | 335.2 KB | edfe940627f6e7c43de8d998cd8de9b5 |
| ENCFF175ACH.bed.gz.dvc | 100.0 B | 6ff8809023dd2ffa63a534b576f327e6 |
| ENCFF175ACH.tabix.bed.gz | 302.43 KB | 784e19f832fea6f851ff9f66713ef430 |
| ENCFF175ACH.tabix.bed.gz.dvc | 106.0 B | a51c0b5e7b03383b935c68598728d8a4 |
| ENCFF175ACH.tabix.bed.gz.tbi | 171.88 KB | f3482a1e1452cc4c45907eee216d01a6 |
| ENCFF175ACH.tabix.bed.gz.tbi.dvc | 110.0 B | 4e6ce667c23e3877ef393922f42d09a7 |
| genomic_resource.yaml | 2.7 KB | 1444fbee619299554529601b372e20e1 |
| genomic_resource_original.yaml | 2.63 KB | 9c79ff4f198699d8bca626ba59ab012a |
| statistics/ |