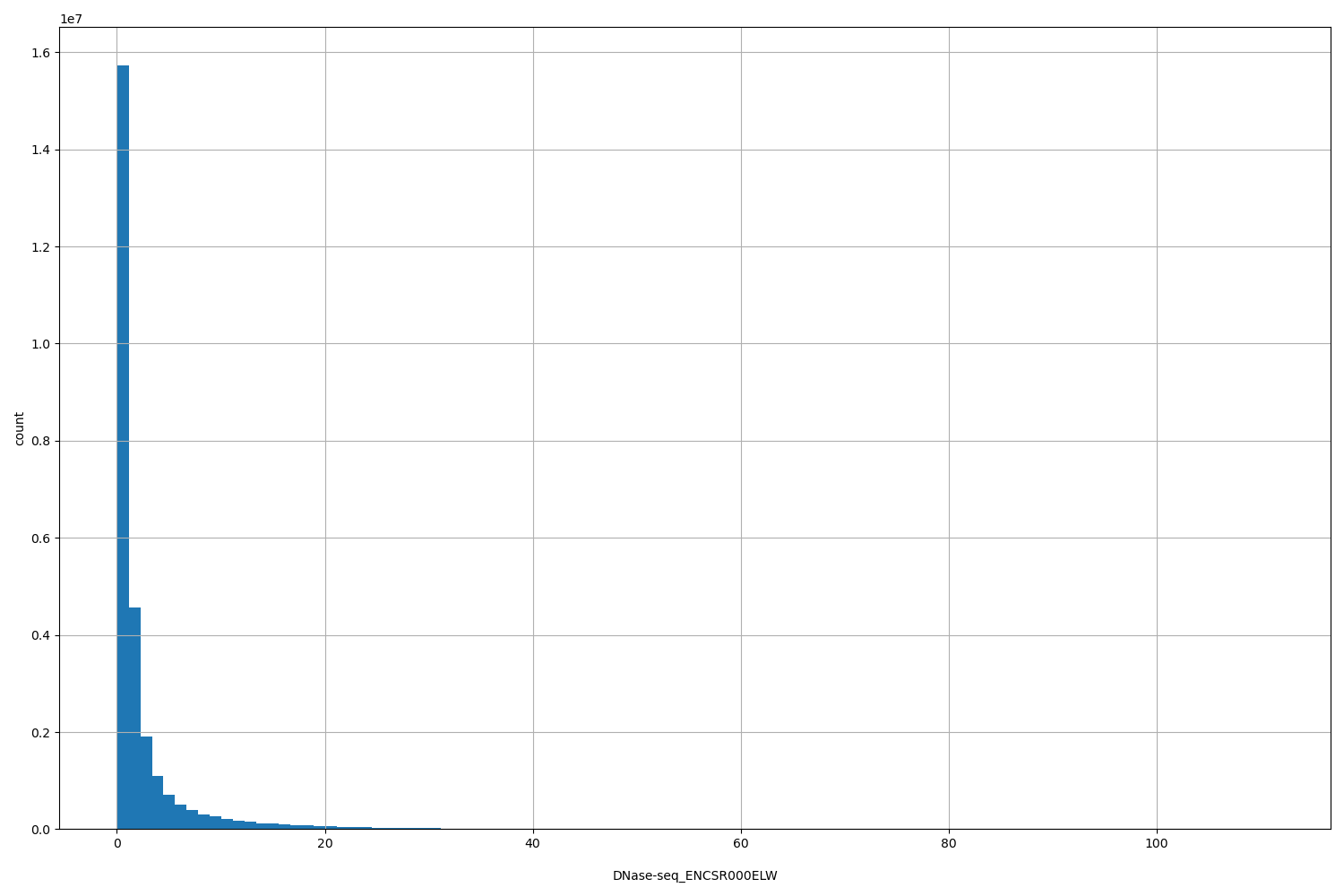
DNase-seq_ENCSR000ELW
| Id: | DNase-seq/ENCSR000ELW |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000ELW [biosample_summary="Homo sapiens A549"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Alignment file {ENCFF194GNS|/files/ENCFF194GNS/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 30796313 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000ELW | float |
DNase-seq_ENCSR000ELW |
DNase-seq ENCSR000ELW [biosample_summary="Homo sapiens A549"]
|
 |
[0.0393, 111] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF128ZVL.bed.gz | 1.55 MB | fe838d13b4f90aee9183969f3f72e4b3 |
| ENCFF128ZVL.bed.gz.dvc | 101.0 B | 5928d3f52aad0da64ee1374e87837059 |
| ENCFF128ZVL.tabix.bed.gz | 1.42 MB | 01ea498d1a6ad6c5bb2ae88b3adc1978 |
| ENCFF128ZVL.tabix.bed.gz.dvc | 107.0 B | 2574108c0dad9b64f7ed154599e2ce1c |
| ENCFF128ZVL.tabix.bed.gz.tbi | 535.55 KB | 399166271864018e7cac287e72f65610 |
| ENCFF128ZVL.tabix.bed.gz.tbi.dvc | 110.0 B | 60828c2da60ac21b364e10a420a5c20b |
| genomic_resource.yaml | 1.69 KB | 72ea1fd174e6db99e3b5fd04eefa31be |
| genomic_resource_original.yaml | 1.63 KB | af241bfc0fe3f319b35ea1f16e90b6c3 |
| statistics/ |