DNase-seq_ENCSR000ELR
| Id: | DNase-seq/ENCSR000ELR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000ELR [biosample_summary="Homo sapiens hepatic stellate cell female adult (59 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (59 years) output_type: peaks audit_error: Alignment file {ENCFF938QTJ|/files/ENCFF938QTJ/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 368818 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_internal_action: Archived analysis {ENCAN808SYS|/analyses/ENCAN808SYS/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF938QTJ|/files/ENCFF938QTJ/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL001DNS|/pipelines/ENCPL001DNS/}) for GRCh38 assembly have a SPOT1 score of 0.26. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
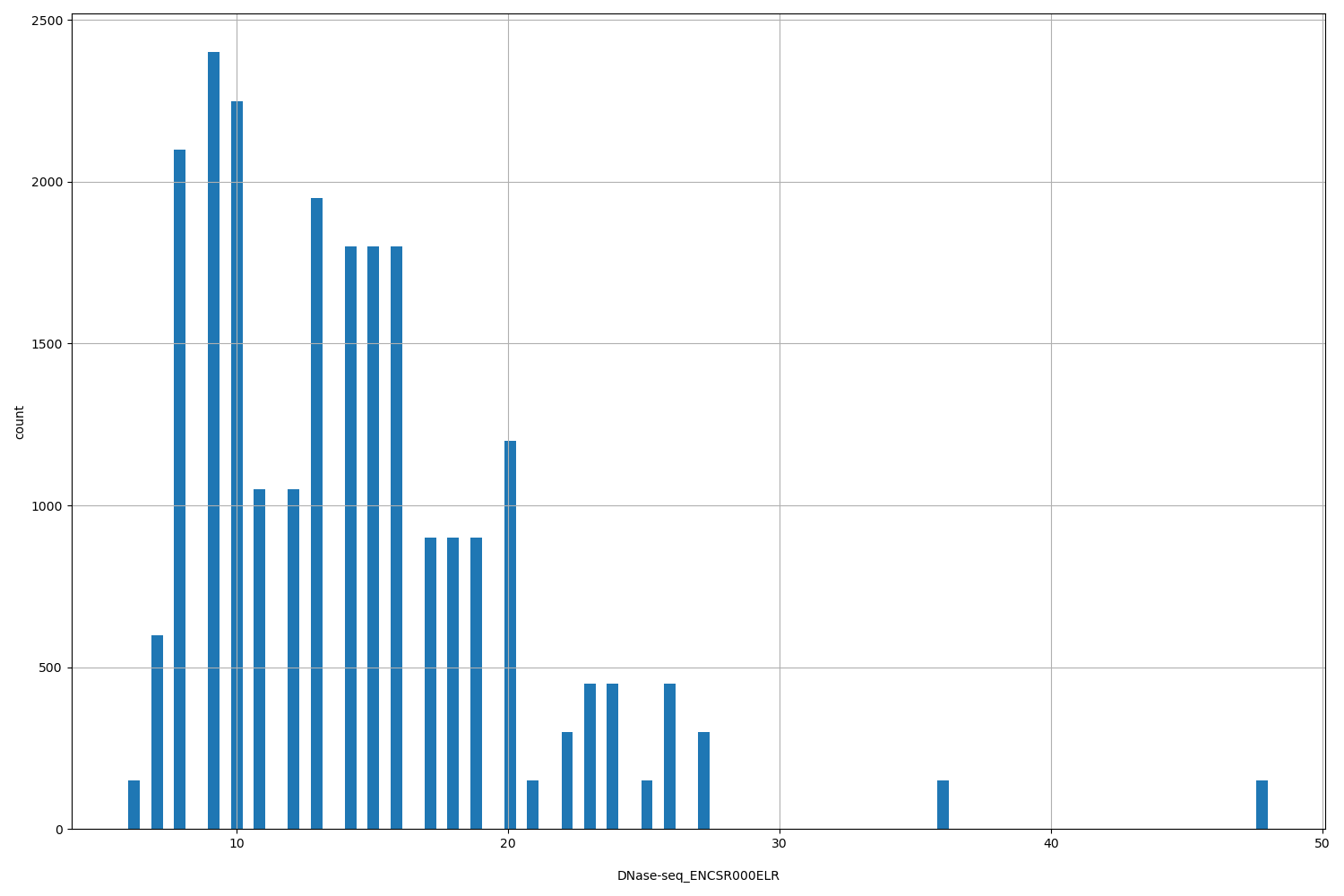
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000ELR | float |
DNase-seq_ENCSR000ELR |
DNase-seq ENCSR000ELR [biosample_summary="Homo sapiens hepatic stellate cell female adult (59 years)"]
|
 |
[6, 48] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF801KLJ.bed.gz | 1.8 KB | b87f0ac776f01076e505d6b6c15c2c46 |
| ENCFF801KLJ.bed.gz.dvc | 98.0 B | f552265dc05265e6b24e0f4b221fc553 |
| ENCFF801KLJ.tabix.bed.gz | 1.49 KB | b2a78583dce4a388a18ed149cf81d8bf |
| ENCFF801KLJ.tabix.bed.gz.dvc | 104.0 B | 05ab5d2042e2db4869622f48f8fe7fda |
| ENCFF801KLJ.tabix.bed.gz.tbi | 5.26 KB | a6a029023d47f0f0262a6022356d3c22 |
| ENCFF801KLJ.tabix.bed.gz.tbi.dvc | 108.0 B | 9e6f9c5f3fca613cad66edeccfee765b |
| genomic_resource.yaml | 2.81 KB | f9d1c6ca6c1ebbc63240958ed209105e |
| genomic_resource_original.yaml | 2.7 KB | 63c78fae968a014e379c5f82e707cd3d |
| statistics/ |