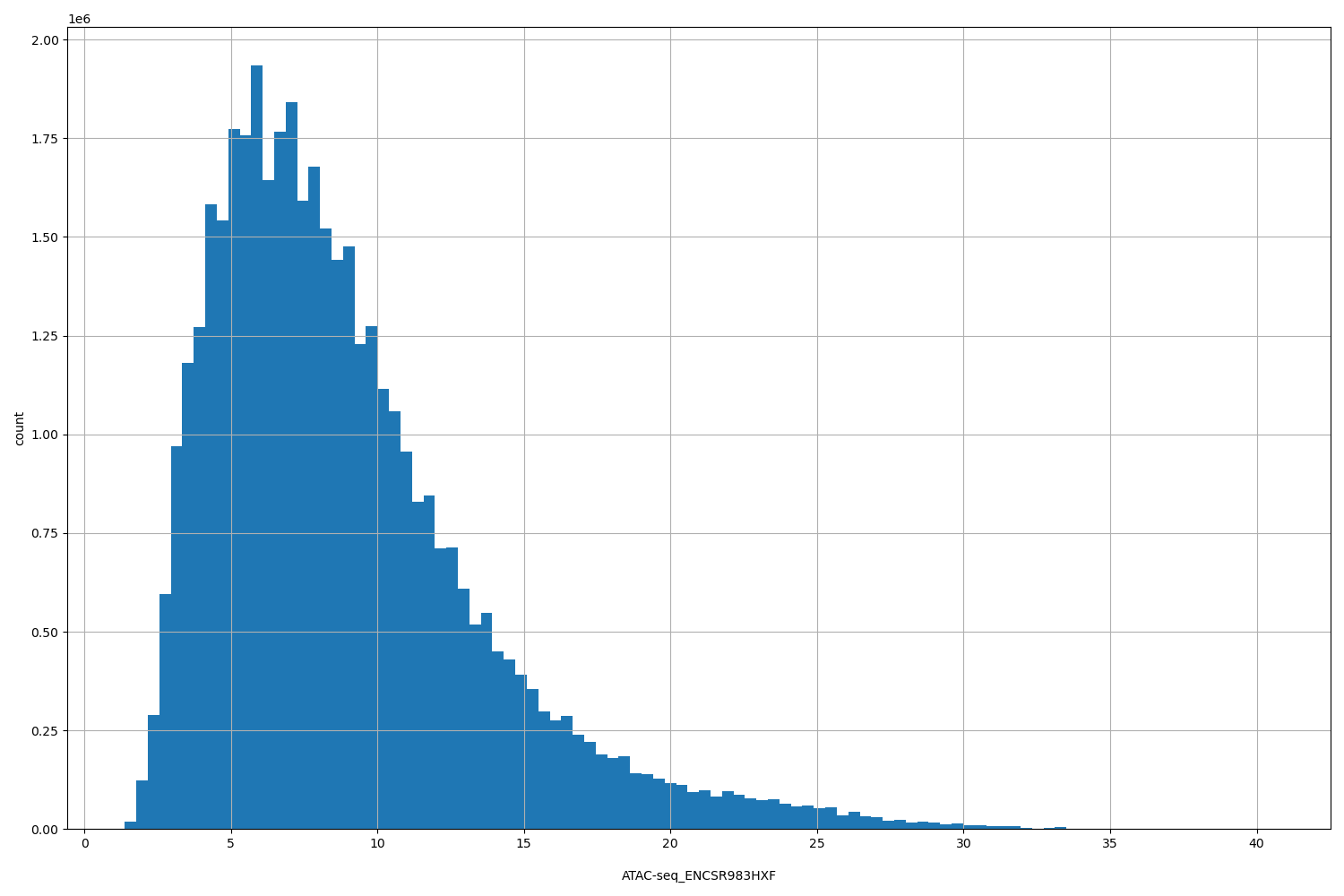
ATAC-seq_ENCSR983HXF
| Id: | ATAC-seq/ENCSR983HXF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR983HXF [biosample_summary="Homo sapiens activated naive CD8-positive, alpha-beta T cell nuclear fraction male adult (30 years) treated with anti-CD3 and anti-CD28 coated beads for 7 days, 10 ng/mL Interleukin-2 for 5 days"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (30 years) treated with anti-CD3 and anti-CD28 coated beads for 7 days, 10 ng/mL Interleukin-2 for 5 days nuclear fraction output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN325KAH|/analyses/ENCAN325KAH/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: Alignment file {ENCFF980SRE|/files/ENCFF980SRE/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline has 23566637 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF980SRE|/files/ENCFF980SRE/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with NRF value of 0.864714. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR983HXF | float |
ATAC-seq_ENCSR983HXF |
ATAC-seq ENCSR983HXF [biosample_summary="Homo sapiens activated naive CD8-positive, alpha-beta T cell nuclear fraction male adult (30 years) treated with anti-CD3 and anti-CD28 coated beads for 7 days, 10 ng/mL Interleukin-2 for 5 days"]
|
 |
[1.38, 40.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF792EFB.bed.gz | 2.96 MB | b0880c8df275fd697862ece62efd8b8b |
| ENCFF792EFB.bed.gz.dvc | 101.0 B | 718f2e87a8717392913bc4ad5e2eb248 |
| ENCFF792EFB.tabix.bed.gz | 806.71 KB | 03ffb5a8da77155d792c2d05961f765e |
| ENCFF792EFB.tabix.bed.gz.dvc | 106.0 B | f3dbee5637a918bd256fe94e60b91e50 |
| ENCFF792EFB.tabix.bed.gz.tbi | 318.94 KB | ad0ec89097ae0f830e2225e29f5e82c8 |
| ENCFF792EFB.tabix.bed.gz.tbi.dvc | 110.0 B | 9231244282ca996a50682e9a0e2c03c2 |
| genomic_resource.yaml | 2.95 KB | 464db00f961031fe567abe9628001833 |
| genomic_resource_original.yaml | 2.72 KB | ce53bd2080db54d368d61f9ebe0fb90c |
| statistics/ |