ATAC-seq_ENCSR956DNB
| Id: | ATAC-seq/ENCSR956DNB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR956DNB [biosample_summary="Homo sapiens K562"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_error: Alignment file {ENCFF445RUW|/files/ENCFF445RUW/} processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline has 8385847 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_error: Alignment file {ENCFF778GYX|/files/ENCFF778GYX/} processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline has 13910063 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_internal_action: Released analysis {ENCAN222XPD|/analyses/ENCAN222XPD/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF842UZU|/files/ENCFF842UZU/} (overlap peaks) and {ENCFF489GQF|/files/ENCFF489GQF/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline have 123697 and 69819 peaks. |
| Labels: |
|
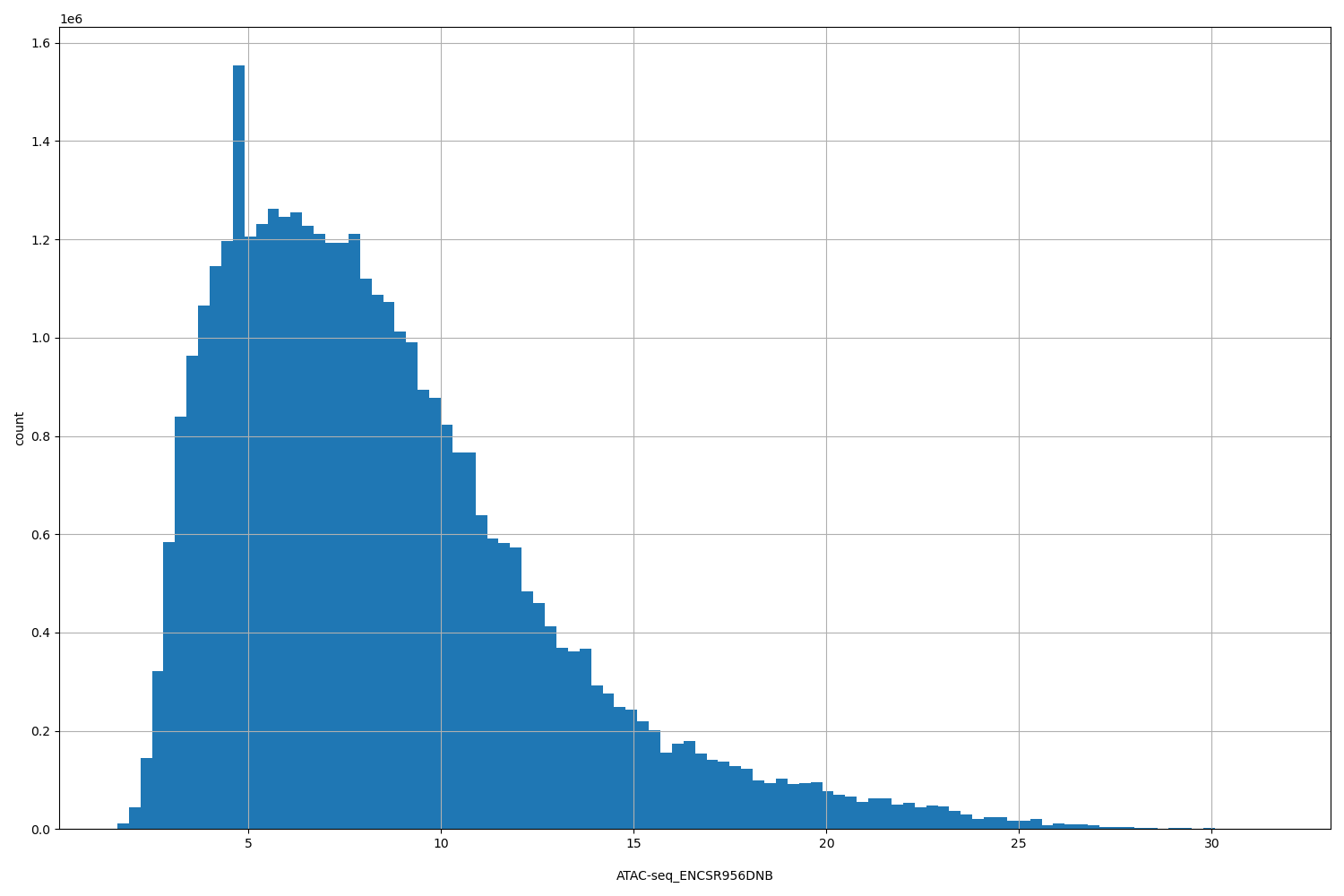
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR956DNB | float |
ATAC-seq_ENCSR956DNB |
ATAC-seq ENCSR956DNB [biosample_summary="Homo sapiens K562"]
|
 |
[1.59, 31.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF842UZU.bed.gz | 2.96 MB | c1738f40f77a19d1773771b87826dbb8 |
| ENCFF842UZU.bed.gz.dvc | 101.0 B | c4eb7e4bf0f5ec047a03f45eec622425 |
| ENCFF842UZU.tabix.bed.gz | 975.34 KB | 74ced2e5939e4951c6320f203acff8fc |
| ENCFF842UZU.tabix.bed.gz.dvc | 106.0 B | 55e03559f7cd36b5bc2300b048219fb6 |
| ENCFF842UZU.tabix.bed.gz.tbi | 389.63 KB | 4b661be2dbf48076b558fedbc35bc0f8 |
| ENCFF842UZU.tabix.bed.gz.tbi.dvc | 110.0 B | 407bbc2dd56695cd45c3ad03bf6d6ca9 |
| genomic_resource.yaml | 2.59 KB | 1d3bdbdd352e16932513ab0bd4749a82 |
| genomic_resource_original.yaml | 2.53 KB | 7a0a11019f754c8d30b38efa100bd005 |
| statistics/ |