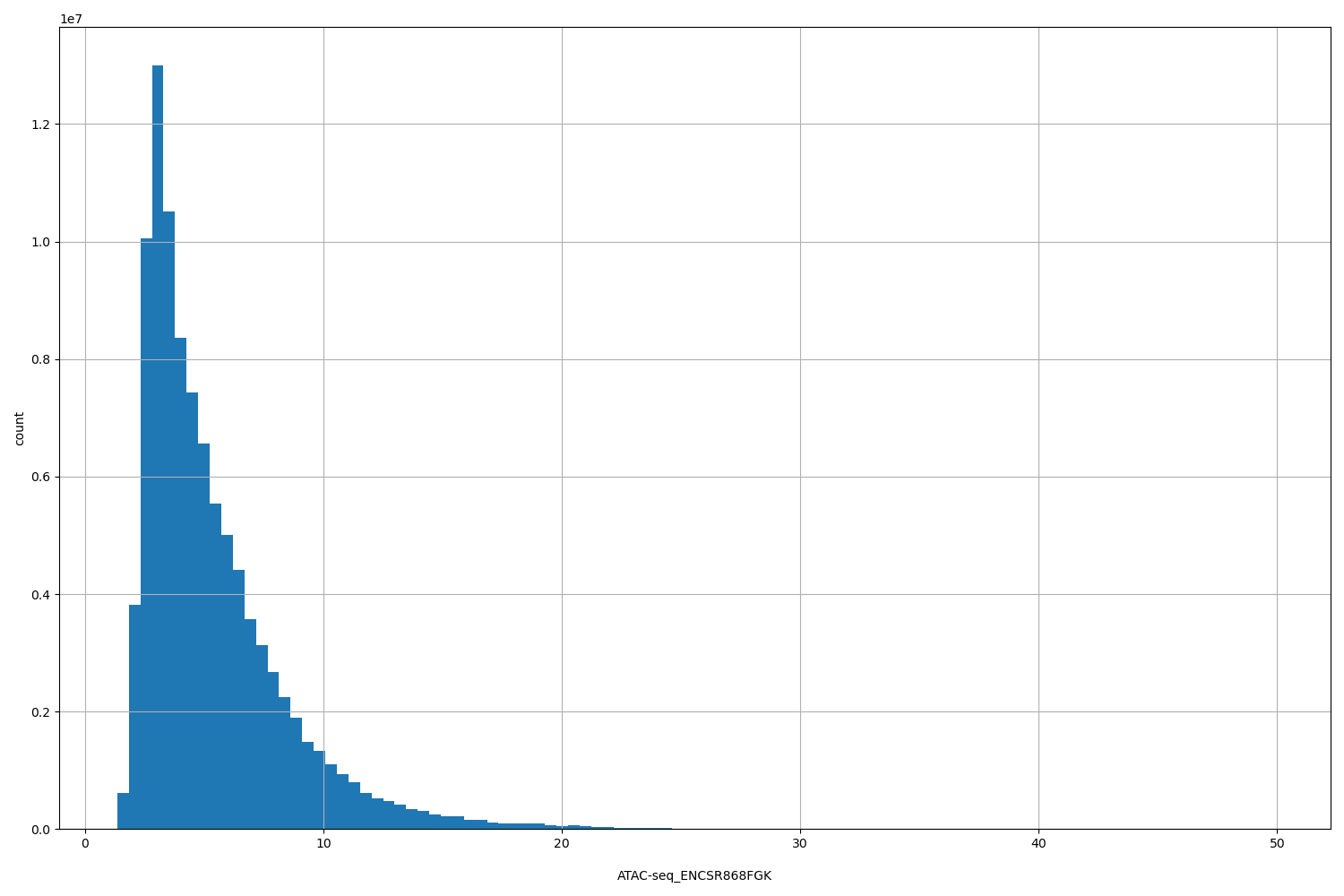
ATAC-seq_ENCSR868FGK
| Id: | ATAC-seq/ENCSR868FGK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR868FGK [biosample_summary="Homo sapiens K562"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN824UKX|/analyses/ENCAN824UKX/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF534DCE|/files/ENCFF534DCE/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.889353. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR868FGK | float |
ATAC-seq_ENCSR868FGK |
ATAC-seq ENCSR868FGK [biosample_summary="Homo sapiens K562"]
|
 |
[1.35, 49.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF333TAT.bed.gz | 6.74 MB | 0f7a6c13e23c2e3fc8716153a89ed481 |
| ENCFF333TAT.bed.gz.dvc | 101.0 B | cca4938c79ffe187ad0f3e20214c42ed |
| ENCFF333TAT.tabix.bed.gz | 1.79 MB | 7ea5b1fb59ae1cf530f4616e86ae2518 |
| ENCFF333TAT.tabix.bed.gz.dvc | 107.0 B | e60c264346d522bf27ae69677f30a3f7 |
| ENCFF333TAT.tabix.bed.gz.tbi | 459.7 KB | 1eaea5b4cfd57d56b336fcf2a6e5359e |
| ENCFF333TAT.tabix.bed.gz.tbi.dvc | 110.0 B | cb2e9e7a469f029aaba7eaab4cc2473a |
| genomic_resource.yaml | 1.82 KB | d233511e54bb6c0d280e39354269d2e1 |
| genomic_resource_original.yaml | 1.75 KB | 8a9cb419caf4efa0a0a81c9374fee2e5 |
| statistics/ |