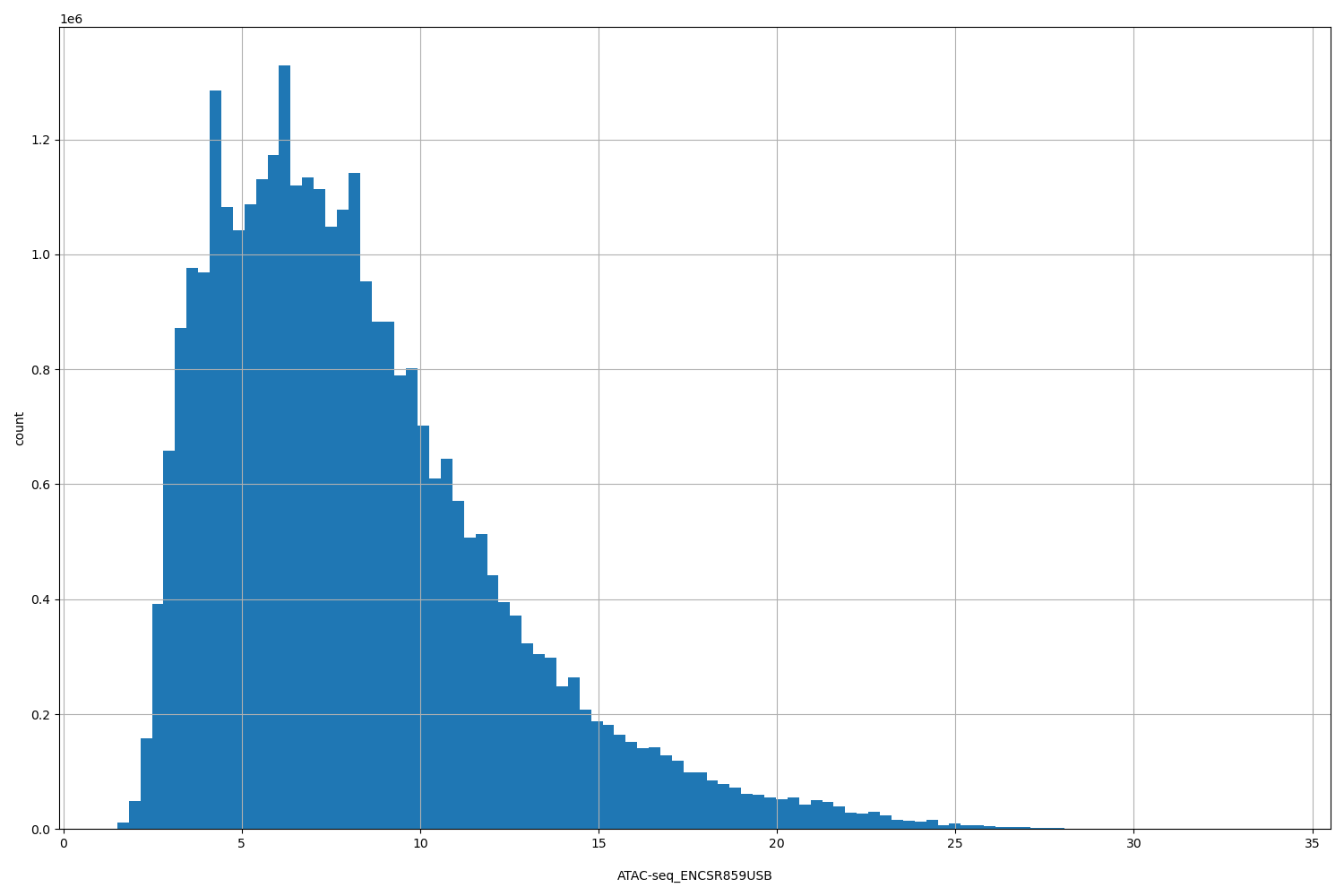
ATAC-seq_ENCSR859USB
| Id: | ATAC-seq/ENCSR859USB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR859USB [biosample_summary="Homo sapiens K562"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_error: Alignment file {ENCFF557RSA|/files/ENCFF557RSA/} processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline has 8585759 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_internal_action: Released analysis {ENCAN615CMZ|/analyses/ENCAN615CMZ/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_not_compliant: Alignment file {ENCFF852RRY|/files/ENCFF852RRY/} processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline has 15065125 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_warning: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF055NNT|/files/ENCFF055NNT/} (overlap peaks) and {ENCFF926KTI|/files/ENCFF926KTI/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v2.1.1 GRCh38 pipeline have 99862 and 55575 peaks. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR859USB | float |
ATAC-seq_ENCSR859USB |
ATAC-seq ENCSR859USB [biosample_summary="Homo sapiens K562"]
|
 |
[1.51, 33.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF055NNT.bed.gz | 2.44 MB | 7f7a4fea9902b4de27656b78a71cdda0 |
| ENCFF055NNT.bed.gz.dvc | 101.0 B | 2f2aa6d1d966a4934b158bea00e74e64 |
| ENCFF055NNT.tabix.bed.gz | 800.17 KB | 5b9abd5100d096de968efd9880e0d7ed |
| ENCFF055NNT.tabix.bed.gz.dvc | 106.0 B | edb6760cbb6f823d11930659cd9b0776 |
| ENCFF055NNT.tabix.bed.gz.tbi | 352.43 KB | cb942756423d44e21d1971ac2b06892e |
| ENCFF055NNT.tabix.bed.gz.tbi.dvc | 110.0 B | 48599476026fc9d4f967e8af22e0eaa7 |
| genomic_resource.yaml | 2.59 KB | aeae9f51ccf0e1f0d02a5820067b891f |
| genomic_resource_original.yaml | 2.52 KB | 9ee9bb22500fb2dcb15f8733ac5dbd7f |
| statistics/ |