ATAC-seq_ENCSR858GTE
| Id: | ATAC-seq/ENCSR858GTE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR858GTE [biosample_summary="Homo sapiens HG03442"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN525YPO|/analyses/ENCAN525YPO/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF130DND|/files/ENCFF130DND/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.755822. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF130DND|/files/ENCFF130DND/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with a PBC1 value of 0.87. |
| Labels: |
|
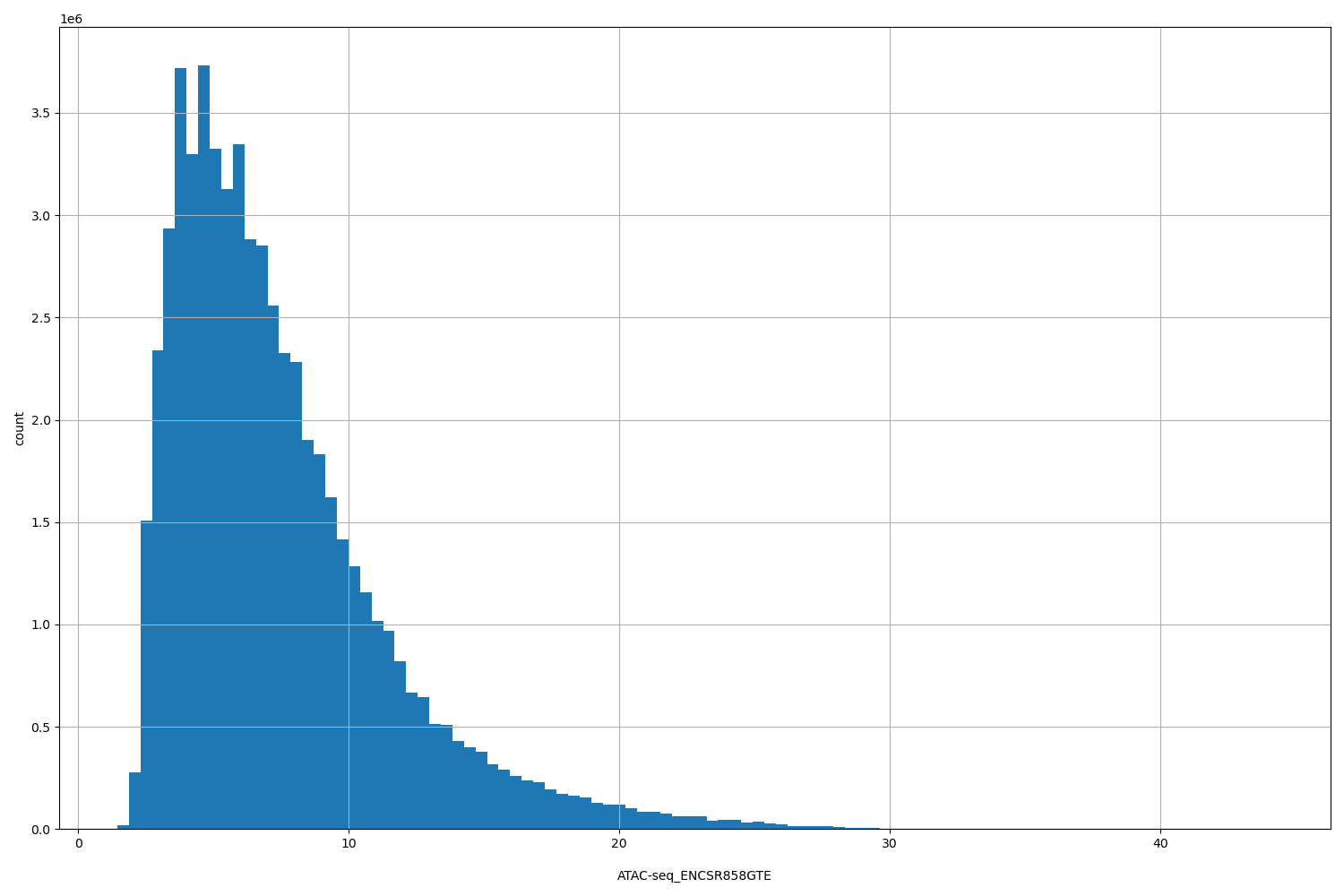
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR858GTE | float |
ATAC-seq_ENCSR858GTE |
ATAC-seq ENCSR858GTE [biosample_summary="Homo sapiens HG03442"]
|
 |
[1.45, 44.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF474JMH.bed.gz | 4.12 MB | c61d390cc11c15ee9c65767b39b37699 |
| ENCFF474JMH.bed.gz.dvc | 101.0 B | 2b3cc797bcf7378d1493dc35501c6f34 |
| ENCFF474JMH.tabix.bed.gz | 1.28 MB | d2f33990c773b31dee9895c3953fc3a9 |
| ENCFF474JMH.tabix.bed.gz.dvc | 107.0 B | 1bf06e8b91f83991228d64ae573ef277 |
| ENCFF474JMH.tabix.bed.gz.tbi | 503.52 KB | 8d7f9fbf827f0b8af21846b9569f56ed |
| ENCFF474JMH.tabix.bed.gz.tbi.dvc | 110.0 B | be52b9938b36002e20ce00aead848e88 |
| genomic_resource.yaml | 2.49 KB | 560ee9af65fbc557679d5c395f1bb4de |
| genomic_resource_original.yaml | 2.42 KB | 62db16bbed3133cf3e92ab4eead11443 |
| statistics/ |