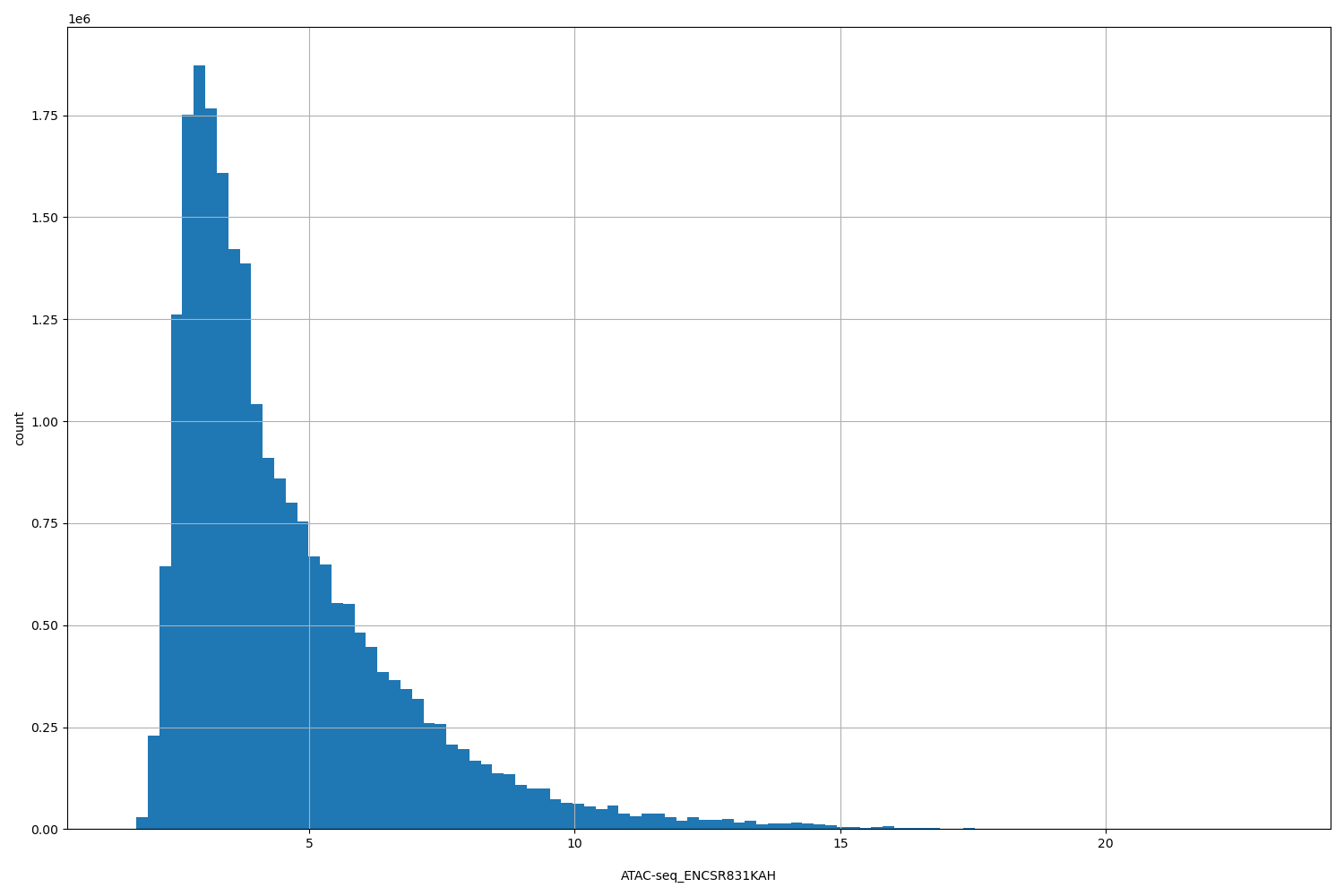
ATAC-seq_ENCSR831KAH
| Id: | ATAC-seq/ENCSR831KAH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR831KAH [biosample_summary="Homo sapiens tibial nerve tissue female adult (51 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (51 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN540JRV|/analyses/ENCAN540JRV/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_internal_action: Released analysis {ENCAN540JRV|/analyses/ENCAN540JRV/} has in progress subobject document {67cd13ee-283c-4db5-b014-84fbdd77acf9|/documents/67cd13ee-283c-4db5-b014-84fbdd77acf9/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF512THA|/files/ENCFF512THA/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.06. audit_not_compliant: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF512THA|/files/ENCFF512THA/} (overlap peaks) and {ENCFF623GOH|/files/ENCFF623GOH/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline have 74075 and 28971 peaks. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR831KAH | float |
ATAC-seq_ENCSR831KAH |
ATAC-seq ENCSR831KAH [biosample_summary="Homo sapiens tibial nerve tissue female adult (51 years)"]
|
 |
[1.53, 23.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF512THA.bed.gz | 1.72 MB | b786d4d249b311903fd7795ac0f99058 |
| ENCFF512THA.bed.gz.dvc | 101.0 B | 80cf62f43a7c9684837a88d7d9e7ef1e |
| ENCFF512THA.tabix.bed.gz | 566.42 KB | ea52a70763ea2bdb2ad0f8a83f9ef35c |
| ENCFF512THA.tabix.bed.gz.dvc | 106.0 B | 37cb4f968f880b0d62b26ccfb3c63553 |
| ENCFF512THA.tabix.bed.gz.tbi | 310.24 KB | c191971bc99d4799f7e751768e39487c |
| ENCFF512THA.tabix.bed.gz.tbi.dvc | 110.0 B | ab9635bed79576f7d954a1b3de3cc34b |
| genomic_resource.yaml | 2.59 KB | a8fe78ab3c040942dc9ef2a255c5caf8 |
| genomic_resource_original.yaml | 2.49 KB | 3759778724e40262917262b5cd5779b7 |
| statistics/ |