ATAC-seq_ENCSR802GEV
| Id: | ATAC-seq/ENCSR802GEV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR802GEV [biosample_summary="Homo sapiens cerebellum tissue male adult (20 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (20 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN059BTS|/analyses/ENCAN059BTS/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF081WNU|/files/ENCFF081WNU/} processed by ATAC-seq ENCODE4 v1.9.2 GRCh38 pipeline has a FRiP score of 0.08. audit_not_compliant: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF081WNU|/files/ENCFF081WNU/} (overlap peaks) and {ENCFF858CCP|/files/ENCFF858CCP/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.9.2 GRCh38 pipeline have 55243 and 26034 peaks. audit_warning: Alignment file {ENCFF111JBV|/files/ENCFF111JBV/} processed by ATAC-seq ENCODE4 v1.9.2 GRCh38 pipeline has 24700277 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF111JBV|/files/ENCFF111JBV/} processed by ATAC-seq ENCODE4 v1.9.2 GRCh38 pipeline was generated from a library with NRF value of 0.84668. |
| Labels: |
|
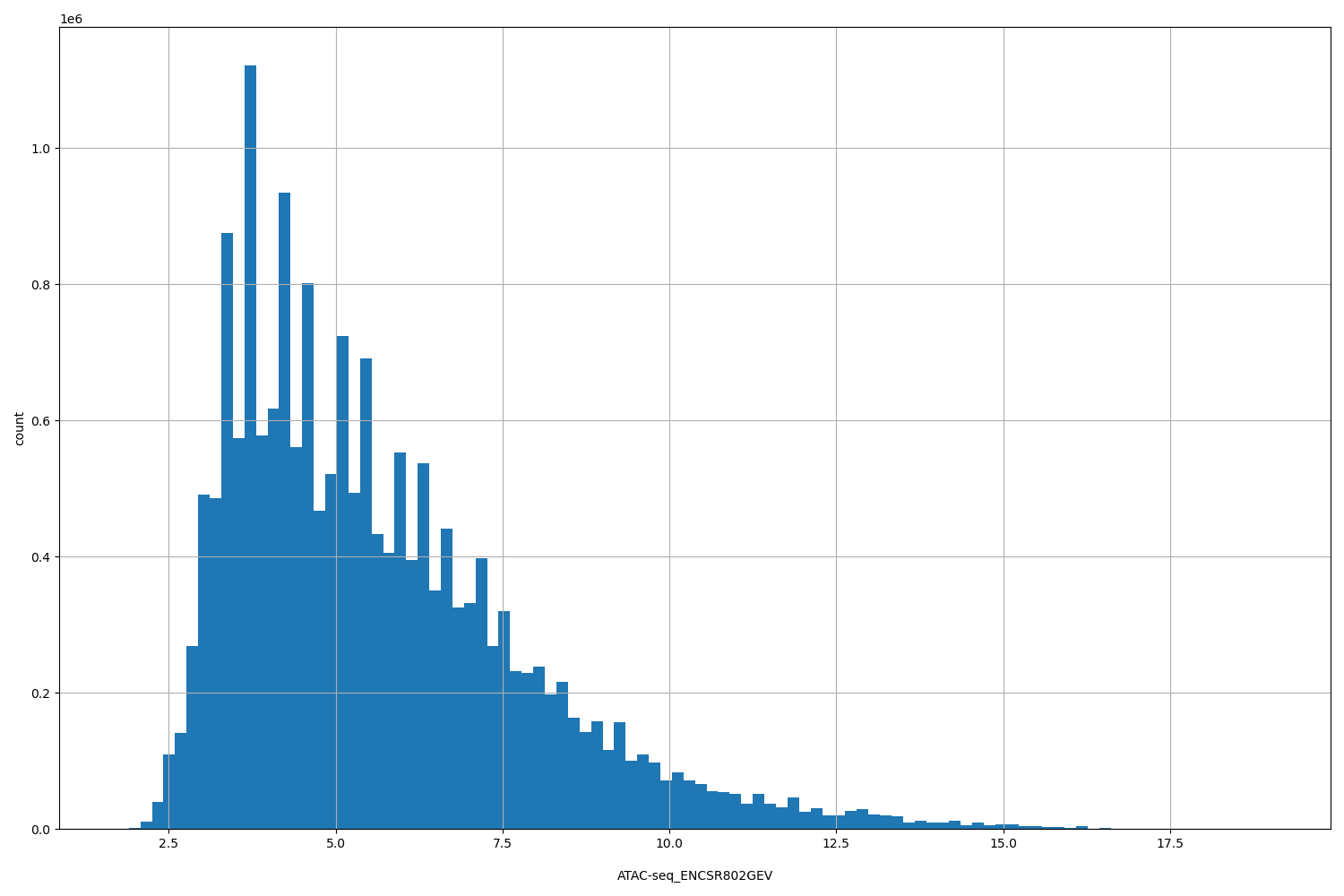
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR802GEV | float |
ATAC-seq_ENCSR802GEV |
ATAC-seq ENCSR802GEV [biosample_summary="Homo sapiens cerebellum tissue male adult (20 years)"]
|
 |
[1.73, 19] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF081WNU.bed.gz | 1.26 MB | 97317bfa6775175703088eeacacdd6b1 |
| ENCFF081WNU.bed.gz.dvc | 101.0 B | 4ef7c98ef5905f416731a329e47b23c7 |
| ENCFF081WNU.tabix.bed.gz | 463.55 KB | c91bc9b1082d6c3c33641991a2696064 |
| ENCFF081WNU.tabix.bed.gz.dvc | 106.0 B | 3b454fe77ad80b998d68bb6b7035e33e |
| ENCFF081WNU.tabix.bed.gz.tbi | 284.08 KB | a7906732b29ba8df8f3fe07c3c685434 |
| ENCFF081WNU.tabix.bed.gz.tbi.dvc | 110.0 B | e9dece22bc5317a4cab8bbb69c9a81fd |
| genomic_resource.yaml | 3.25 KB | c549830dc9631ec1284edc45b3118dde |
| genomic_resource_original.yaml | 3.15 KB | 0d2cad895e054c2026f56d78865af71e |
| statistics/ |