ATAC-seq_ENCSR775KMA
| Id: | ATAC-seq/ENCSR775KMA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR775KMA [biosample_summary="Homo sapiens activated T-cell male adult (38 years) treated with 50 U/mL Interleukin-2 for 4 hours, anti-CD3 and anti-CD28 coated beads for 4 hours"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (38 years) treated with 50 U/mL Interleukin-2 for 4 hours, anti-CD3 and anti-CD28 coated beads for 4 hours output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN243QOP|/analyses/ENCAN243QOP/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: Alignment file {ENCFF880NEB|/files/ENCFF880NEB/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline has 21803135 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. audit_warning: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF089LSF|/files/ENCFF089LSF/} (overlap peaks) and {ENCFF760LQQ|/files/ENCFF760LQQ/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline have 99952 and 63133 peaks. |
| Labels: |
|
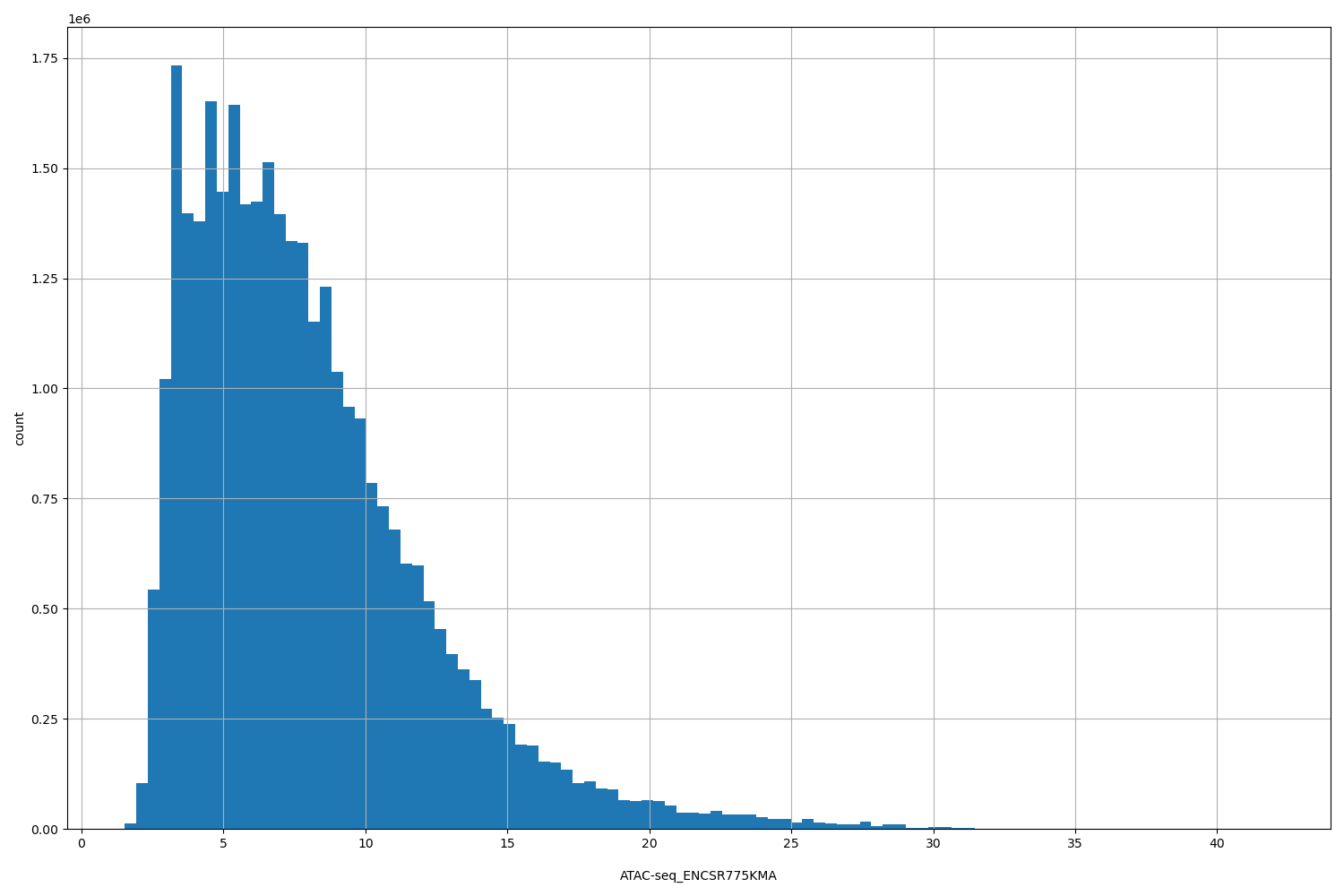
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR775KMA | float |
ATAC-seq_ENCSR775KMA |
ATAC-seq ENCSR775KMA [biosample_summary="Homo sapiens activated T-cell male adult (38 years) treated with 50 U/mL Interleukin-2 for 4 hours, anti-CD3 and anti-CD28 coated beads for 4 hours"]
|
 |
[1.53, 42] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF089LSF.bed.gz | 2.35 MB | dabdc1bc6225436dbf36d43da3d2c1e1 |
| ENCFF089LSF.bed.gz.dvc | 101.0 B | e768ed0ef26c5cc99b36bf77b34e287f |
| ENCFF089LSF.tabix.bed.gz | 644.24 KB | 8b6d834b9d87e45d50c7d6f8b0e6f216 |
| ENCFF089LSF.tabix.bed.gz.dvc | 106.0 B | edc4a51bb7385a016f101112d18f86c4 |
| ENCFF089LSF.tabix.bed.gz.tbi | 297.13 KB | 1f9a15572467306f4f6a96d63b78491e |
| ENCFF089LSF.tabix.bed.gz.tbi.dvc | 110.0 B | 0bb880be30d181f6b7e2ca5056034ebe |
| genomic_resource.yaml | 2.81 KB | 2f9711d7437e252d820280006b27aeaa |
| genomic_resource_original.yaml | 2.62 KB | 46d551226bee40b1dc76d16360e019dc |
| statistics/ |