ATAC-seq_ENCSR705RZX
| Id: | ATAC-seq/ENCSR705RZX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR705RZX [biosample_summary="Homo sapiens uterus tissue female adult (46 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (46 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN485JSN|/analyses/ENCAN485JSN/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF705PUR|/files/ENCFF705PUR/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.07. audit_warning: Transcription Start Site (TSS) enrichment values for alignments to the human genome GRCh38 are concerning when < 5, acceptable between 5 and 7, and ideal when > 7. ENCODE processed file {ENCFF408ZHS|/files/ENCFF408ZHS/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a TSS enrichment value of 6.8988414089210055. audit_warning: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF705PUR|/files/ENCFF705PUR/} (overlap peaks) and {ENCFF344WAS|/files/ENCFF344WAS/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline have 137906 and 56424 peaks. |
| Labels: |
|
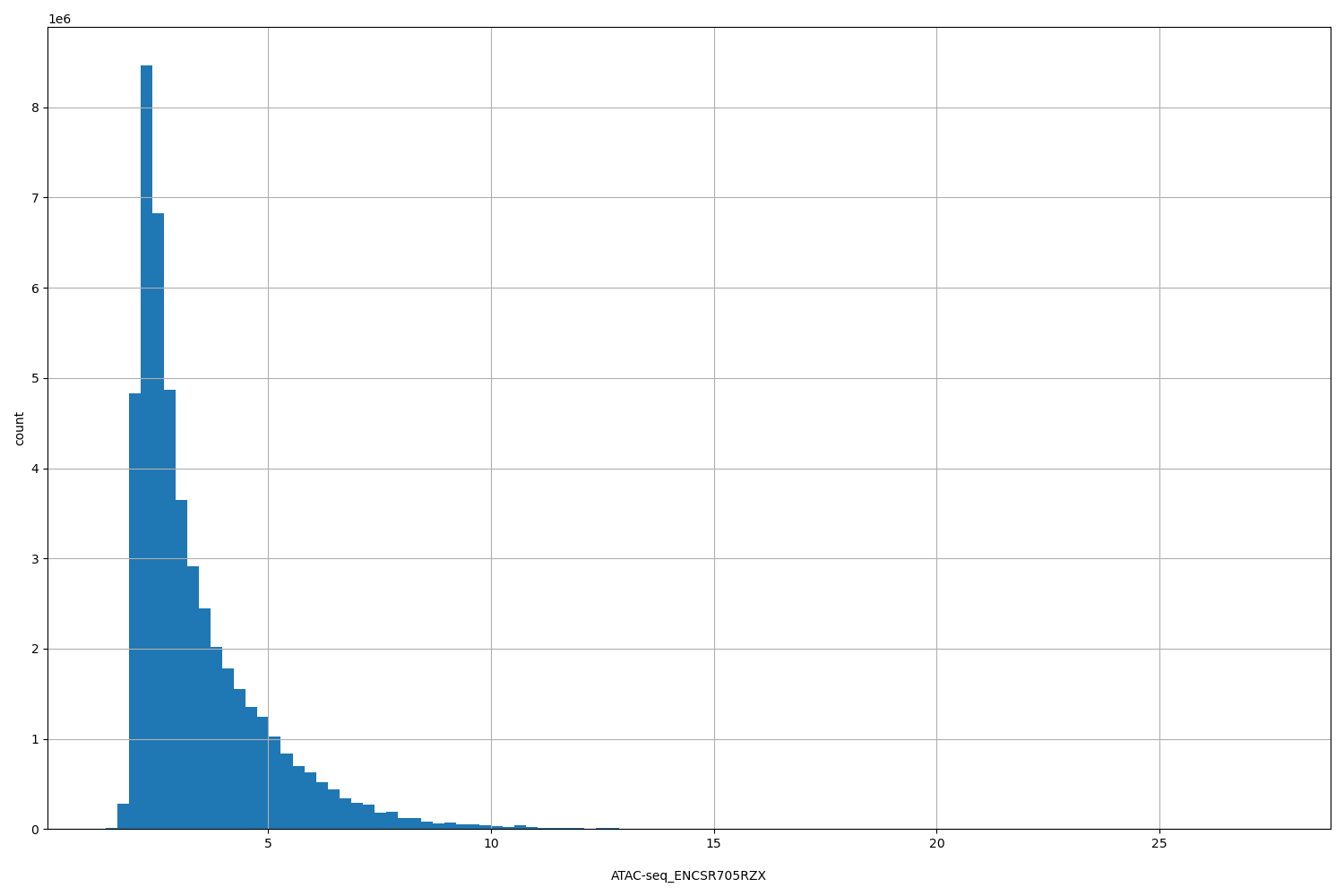
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR705RZX | float |
ATAC-seq_ENCSR705RZX |
ATAC-seq ENCSR705RZX [biosample_summary="Homo sapiens uterus tissue female adult (46 years)"]
|
 |
[1.37, 27.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF705PUR.bed.gz | 3.15 MB | 981dfd4d45dac178b6f605ecfdaa8a80 |
| ENCFF705PUR.bed.gz.dvc | 101.0 B | f879041bf986ea418015a5bb54da6169 |
| ENCFF705PUR.tabix.bed.gz | 984.27 KB | 11b61344e002b8362c397fb4b34446c5 |
| ENCFF705PUR.tabix.bed.gz.dvc | 107.0 B | 170acddb973b8068153770f35d798eef |
| ENCFF705PUR.tabix.bed.gz.tbi | 446.74 KB | 947166c23d9e162b77eb83c3d6f22088 |
| ENCFF705PUR.tabix.bed.gz.tbi.dvc | 110.0 B | c9e33edd4781355eb3014106f453a343 |
| genomic_resource.yaml | 3.07 KB | 0208869870338cc2b20ad447437568c2 |
| genomic_resource_original.yaml | 2.97 KB | d9f0f74f8bb54deb63295490b99cc225 |
| statistics/ |