ATAC-seq_ENCSR608KJD
ATAC-seq ENCSR608KJD [biosample_summary="Homo sapiens HG03097"]
| Id: | ATAC-seq/ENCSR608KJD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR608KJD [biosample_summary="Homo sapiens HG03097"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN924NWJ|/analyses/ENCAN924NWJ/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF720PYC|/files/ENCFF720PYC/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.19. |
| Labels: |
|
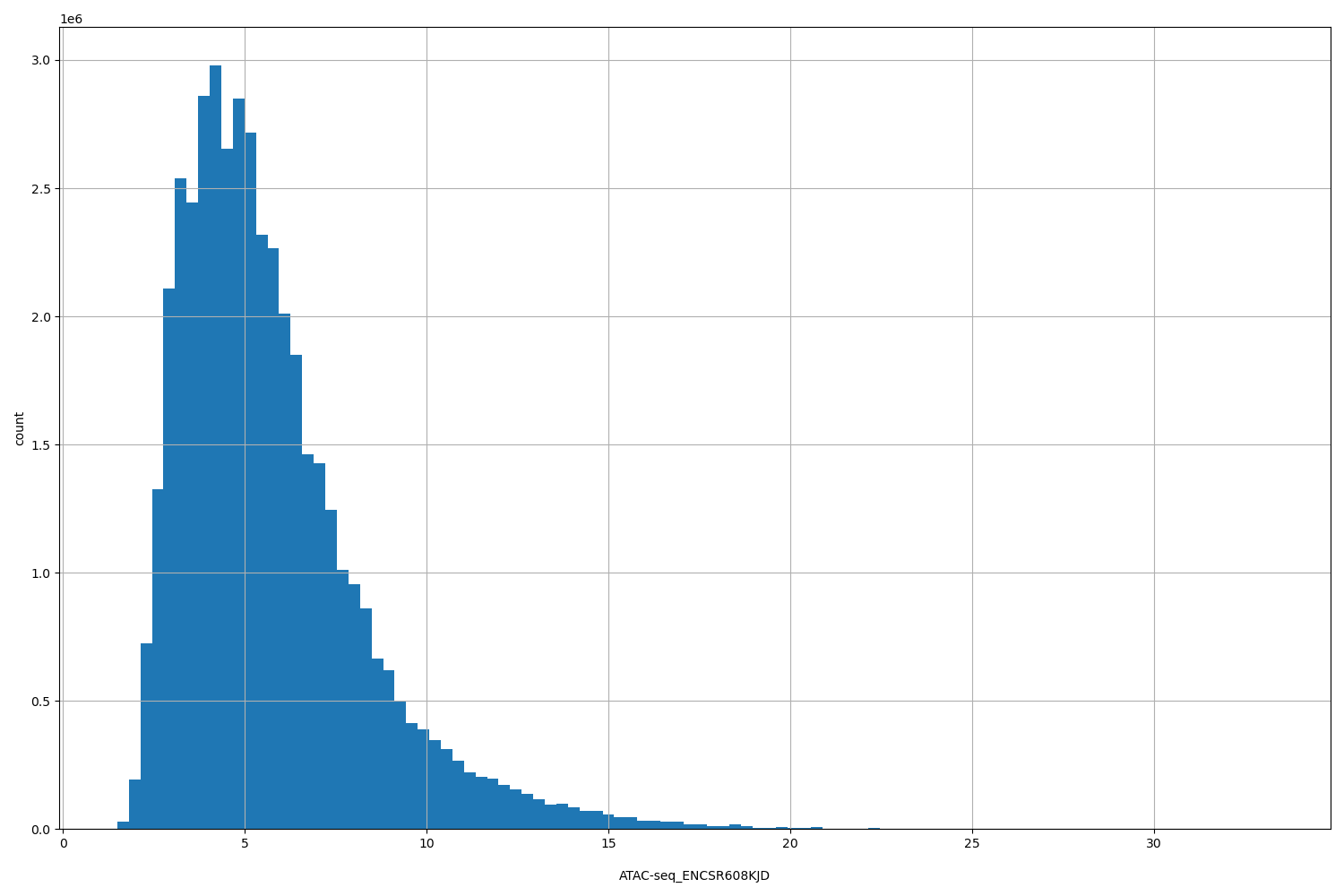
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR608KJD | float |
ATAC-seq_ENCSR608KJD |
ATAC-seq ENCSR608KJD [biosample_summary="Homo sapiens HG03097"]
|
 |
[1.5, 33.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF720PYC.bed.gz | 3.17 MB | 748b8d5a4ea1bc0293d166ca26d26b3a |
| ENCFF720PYC.bed.gz.dvc | 101.0 B | 629b24eb156717d648dcd29559344cc0 |
| ENCFF720PYC.tabix.bed.gz | 959.85 KB | 36130a0db1e6467c9483f67100d97b47 |
| ENCFF720PYC.tabix.bed.gz.dvc | 106.0 B | ed0e136d7913bda877adb2bd8be15ebb |
| ENCFF720PYC.tabix.bed.gz.tbi | 403.03 KB | a29b492265177f8161b523831a7c38d6 |
| ENCFF720PYC.tabix.bed.gz.tbi.dvc | 110.0 B | 617873b050fcddfa4f131d6c857f9194 |
| genomic_resource.yaml | 1.71 KB | 1b06e200b5de6b58a00c33587c84ba15 |
| genomic_resource_original.yaml | 1.64 KB | d21f4420cdc0cc13ecfa16c546954c86 |
| statistics/ |