ATAC-seq_ENCSR591PIX
| Id: | ATAC-seq/ENCSR591PIX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR591PIX [biosample_summary="Homo sapiens Panc1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN507JXY|/analyses/ENCAN507JXY/} has in progress subobject document {ff895ea1-62f2-4f38-9454-a7a15e118e88|/documents/ff895ea1-62f2-4f38-9454-a7a15e118e88/} audit_internal_action: Released analysis {ENCAN507JXY|/analyses/ENCAN507JXY/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF822BKT|/files/ENCFF822BKT/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.875659. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF822BKT|/files/ENCFF822BKT/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with a PBC1 value of 0.89. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF836WDC|/files/ENCFF836WDC/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.857134. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF836WDC|/files/ENCFF836WDC/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with a PBC1 value of 0.89. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR591PIX | float |
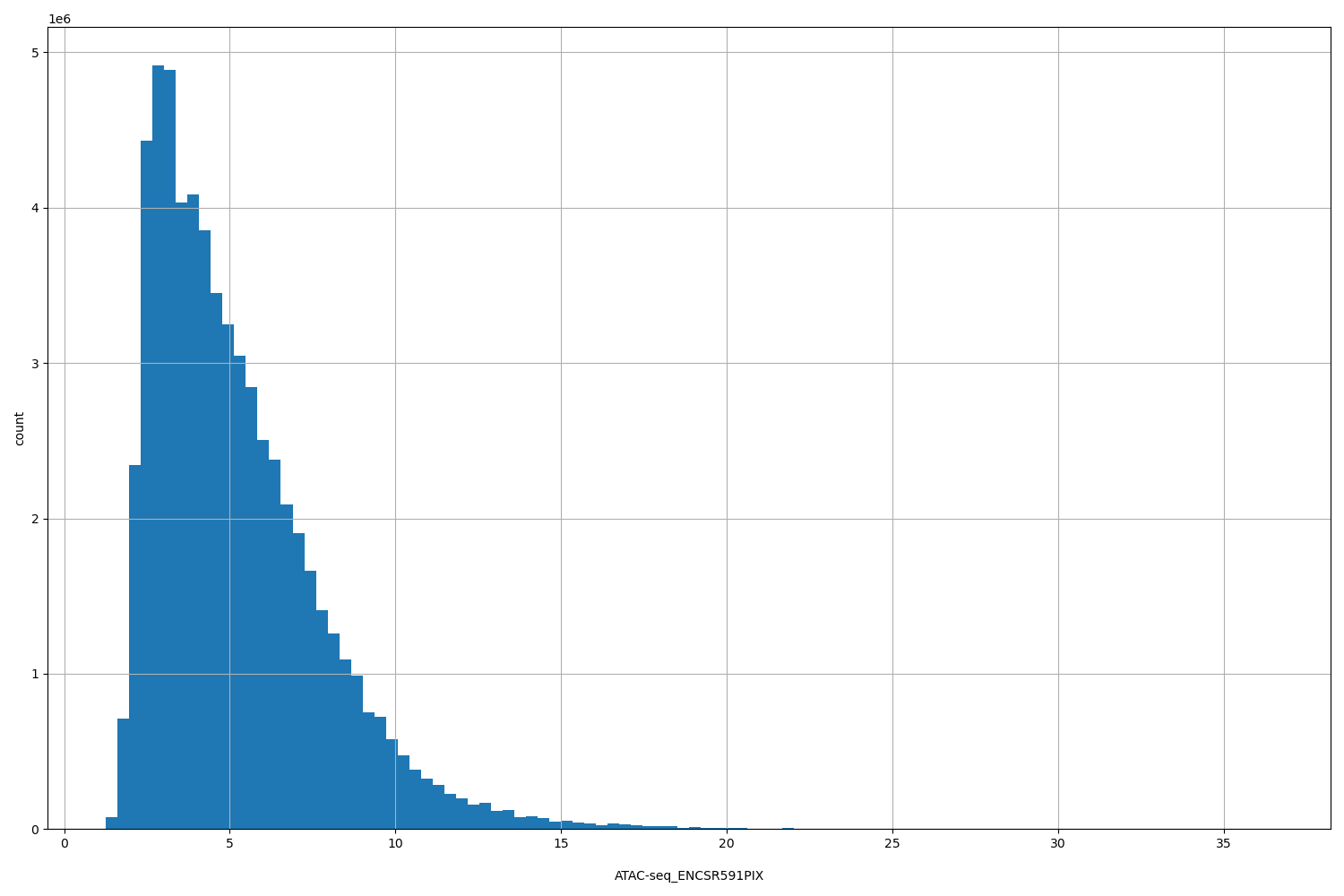
ATAC-seq_ENCSR591PIX |
ATAC-seq ENCSR591PIX [biosample_summary="Homo sapiens Panc1"]
|
 |
[1.26, 36.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF182SSP.bed.gz | 4.97 MB | a3c499c3fb055a084365eaf9ac8bb19f |
| ENCFF182SSP.bed.gz.dvc | 101.0 B | 327653ed60902c3ca28c61313f10da59 |
| ENCFF182SSP.tabix.bed.gz | 1.15 MB | 87d3ce848e8f87b83f9ef72d50ab3c0a |
| ENCFF182SSP.tabix.bed.gz.dvc | 107.0 B | bdd2e92de1420ea0df4b1eaa56845533 |
| ENCFF182SSP.tabix.bed.gz.tbi | 387.22 KB | c5e50de6d605739b44f66b18b9e9e4d7 |
| ENCFF182SSP.tabix.bed.gz.tbi.dvc | 110.0 B | 7a615555f0f4736ab38cbd94ff2d031a |
| genomic_resource.yaml | 3.9 KB | 32acd1cbd4edbd14087933710864cc06 |
| genomic_resource_original.yaml | 3.83 KB | fcf097bfdd94cfacfe409cd5418125ad |
| statistics/ |