ATAC-seq_ENCSR584AXZ
| Id: | ATAC-seq/ENCSR584AXZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR584AXZ [biosample_summary="Homo sapiens coronary artery tissue female adult (51 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (51 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN960SLX|/analyses/ENCAN960SLX/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_internal_action: Released analysis {ENCAN960SLX|/analyses/ENCAN960SLX/} has in progress subobject document {d3be7c62-5279-4d3e-b698-86c1ca80c17d|/documents/d3be7c62-5279-4d3e-b698-86c1ca80c17d/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF442OCV|/files/ENCFF442OCV/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.07. audit_not_compliant: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF442OCV|/files/ENCFF442OCV/} (overlap peaks) and {ENCFF231FCU|/files/ENCFF231FCU/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline have 82924 and 36671 peaks. |
| Labels: |
|
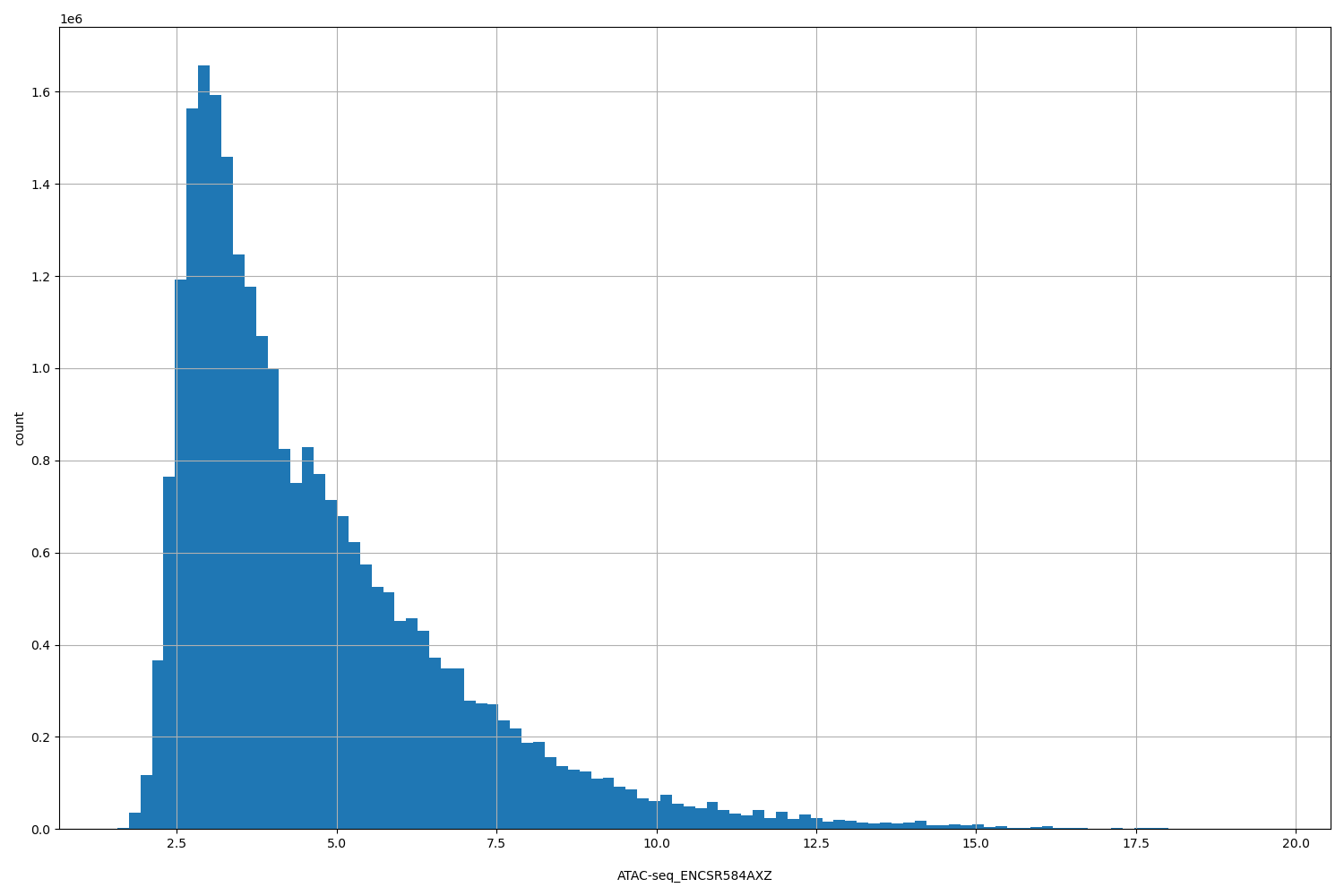
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR584AXZ | float |
ATAC-seq_ENCSR584AXZ |
ATAC-seq ENCSR584AXZ [biosample_summary="Homo sapiens coronary artery tissue female adult (51 years)"]
|
 |
[1.57, 19.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF442OCV.bed.gz | 1.94 MB | 813f09d6a7d464b282ce3e98245b9b6a |
| ENCFF442OCV.bed.gz.dvc | 101.0 B | 8352e7506ebd7cccb5160a6809a0a73c |
| ENCFF442OCV.tabix.bed.gz | 631.36 KB | c390bcdb9f8a7b421dbcd46887b3ed2f |
| ENCFF442OCV.tabix.bed.gz.dvc | 106.0 B | c0abbb080365336f6fbfc733d5380d8a |
| ENCFF442OCV.tabix.bed.gz.tbi | 330.96 KB | cb9ff3aa86f596017109f75d74ed09bf |
| ENCFF442OCV.tabix.bed.gz.tbi.dvc | 110.0 B | 3dabb5755deebaf5403a26f46e93ffbf |
| genomic_resource.yaml | 2.6 KB | 73b2b28bae037fe31d79782240c34c2c |
| genomic_resource_original.yaml | 2.5 KB | e100992dbaa29ce374a772b4a8eeec56 |
| statistics/ |