ATAC-seq_ENCSR513EVP
| Id: | ATAC-seq/ENCSR513EVP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR513EVP [biosample_summary="Homo sapiens naive thymus-derived CD8-positive, alpha-beta T cell male adult (42 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (42 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN372DPK|/analyses/ENCAN372DPK/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF036OKE|/files/ENCFF036OKE/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline has a FRiP score of 0.20. audit_not_compliant: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF036OKE|/files/ENCFF036OKE/} (overlap peaks) and {ENCFF242BPO|/files/ENCFF242BPO/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline have 76910 and 43467 peaks. audit_warning: Alignment file {ENCFF858GSU|/files/ENCFF858GSU/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline has 23191739 usable fragments. According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have > 25 million usable fragments. 20-25 million is acceptable and < 15 million is not compliant. |
| Labels: |
|
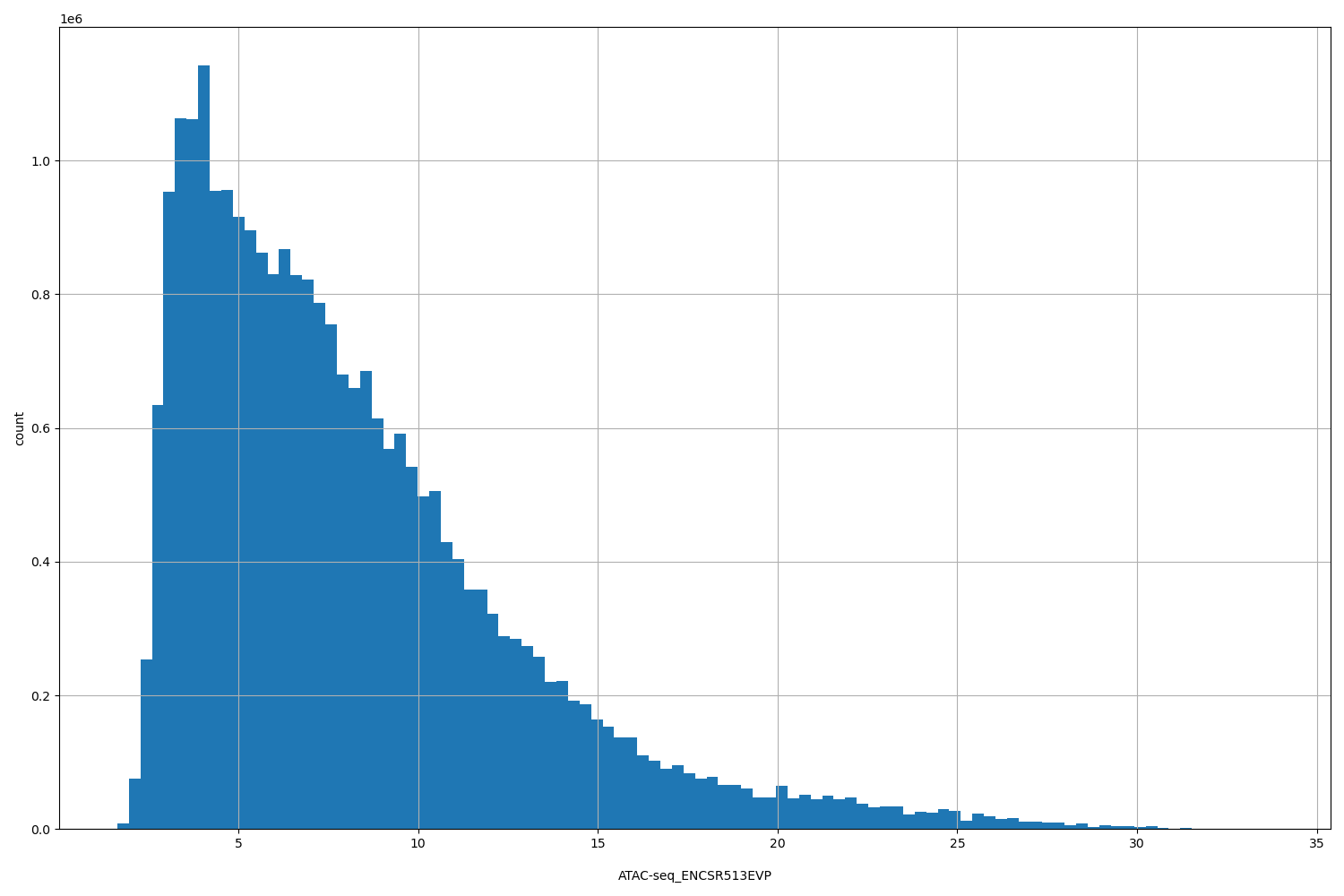
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR513EVP | float |
ATAC-seq_ENCSR513EVP |
ATAC-seq ENCSR513EVP [biosample_summary="Homo sapiens naive thymus-derived CD8-positive, alpha-beta T cell male adult (42 years)"]
|
 |
[1.63, 33.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF036OKE.bed.gz | 1.89 MB | 60f20d33a48fd101271e573bff3cf77b |
| ENCFF036OKE.bed.gz.dvc | 101.0 B | aac7db8ecdc4800be0b650c468abcbed |
| ENCFF036OKE.tabix.bed.gz | 585.1 KB | 3aa3393a41b044cd48e3cb73a211437e |
| ENCFF036OKE.tabix.bed.gz.dvc | 106.0 B | b755edafae91c505ed9081b19c05a25e |
| ENCFF036OKE.tabix.bed.gz.tbi | 278.93 KB | a6b84da4187c52bd0bc40c332978822b |
| ENCFF036OKE.tabix.bed.gz.tbi.dvc | 110.0 B | 38e77081ece04f3e84946f811b121cb8 |
| genomic_resource.yaml | 2.9 KB | 975a5d176f3b440612fcf74aaf674859 |
| genomic_resource_original.yaml | 2.77 KB | 903d3ca4a75b0ed7e6d36d258ab6c264 |
| statistics/ |