ATAC-seq_ENCSR386HAZ
| Id: | ATAC-seq/ENCSR386HAZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR386HAZ [biosample_summary="Homo sapiens transverse colon tissue female adult (51 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (51 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN571WPB|/analyses/ENCAN571WPB/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_internal_action: Released analysis {ENCAN571WPB|/analyses/ENCAN571WPB/} has in progress subobject document {d0d3b944-6c96-48db-8cb4-30011c65929d|/documents/d0d3b944-6c96-48db-8cb4-30011c65929d/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF355GMG|/files/ENCFF355GMG/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.15. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF422XWI|/files/ENCFF422XWI/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.848744. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF422XWI|/files/ENCFF422XWI/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with a PBC1 value of 0.86. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR386HAZ | float |
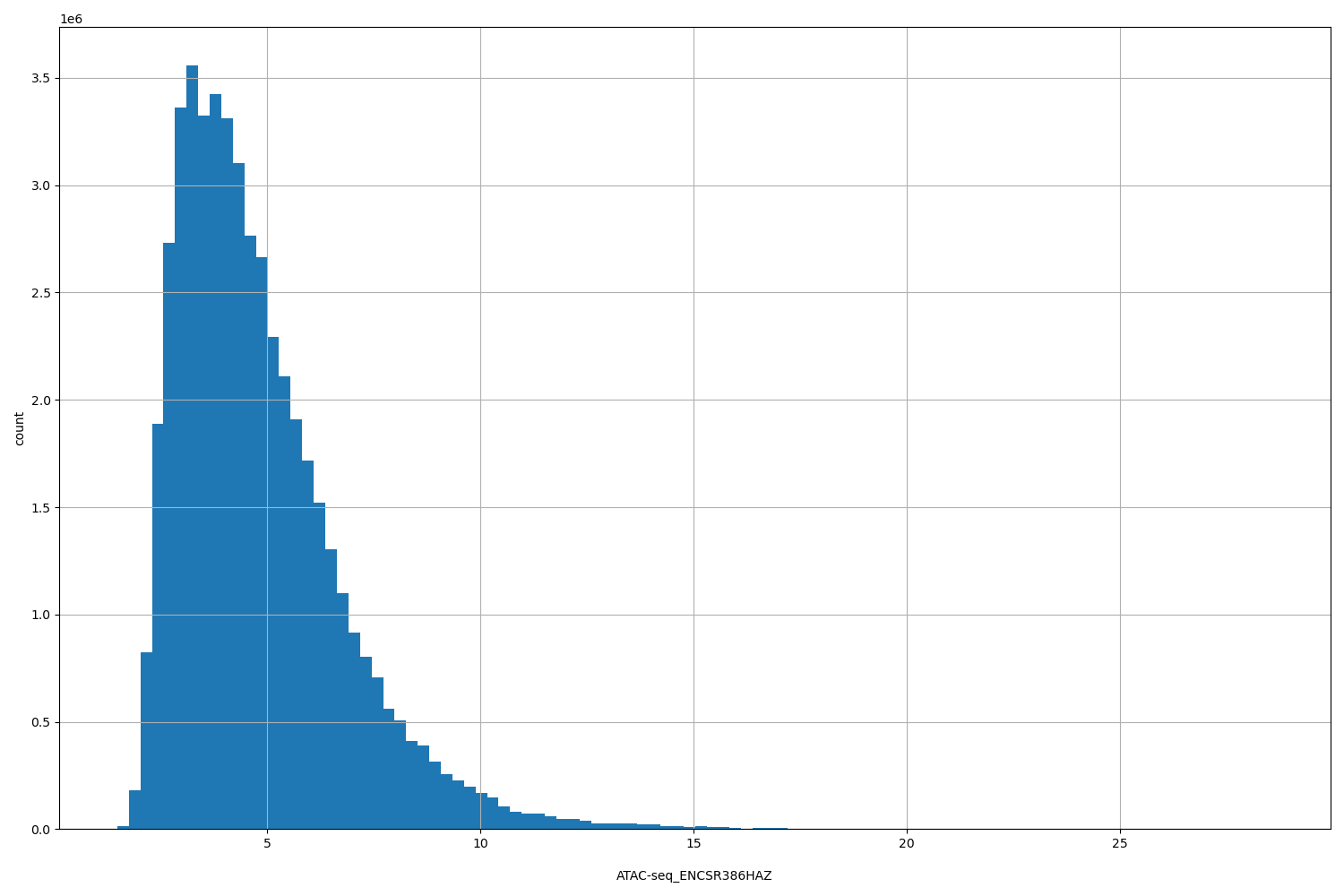
ATAC-seq_ENCSR386HAZ |
ATAC-seq ENCSR386HAZ [biosample_summary="Homo sapiens transverse colon tissue female adult (51 years)"]
|
 |
[1.48, 28.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF355GMG.bed.gz | 3.77 MB | d61e3f1a4aba9927abcda5fd53ba9211 |
| ENCFF355GMG.bed.gz.dvc | 101.0 B | b9896945b9e38e740614617b364bc4c5 |
| ENCFF355GMG.tabix.bed.gz | 1.01 MB | c9b903a7b7421c8bba37e2f0bac0144e |
| ENCFF355GMG.tabix.bed.gz.dvc | 107.0 B | 830359a45971dd0d4c34d201f617610d |
| ENCFF355GMG.tabix.bed.gz.tbi | 407.73 KB | 6cb5d7878c201ed8d99011a617c98f29 |
| ENCFF355GMG.tabix.bed.gz.tbi.dvc | 110.0 B | faad270320abeeea616c4429ba76de36 |
| genomic_resource.yaml | 3.21 KB | 90d8e5703c07259ec68fd8a91df37574 |
| genomic_resource_original.yaml | 3.1 KB | 9544d9515fda64666346f278485c3856 |
| statistics/ |