ATAC-seq_ENCSR376YMU
| Id: | ATAC-seq/ENCSR376YMU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR376YMU [biosample_summary="Homo sapiens HG03439"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN593KEC|/analyses/ENCAN593KEC/} has in progress subobject document {e7fa7f12-9fec-45dc-a526-b595554d5069|/documents/e7fa7f12-9fec-45dc-a526-b595554d5069/} audit_internal_action: Released analysis {ENCAN593KEC|/analyses/ENCAN593KEC/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF326QXM|/files/ENCFF326QXM/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with NRF value of 0.888688. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF326QXM|/files/ENCFF326QXM/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline was generated from a library with a PBC1 value of 0.90. |
| Labels: |
|
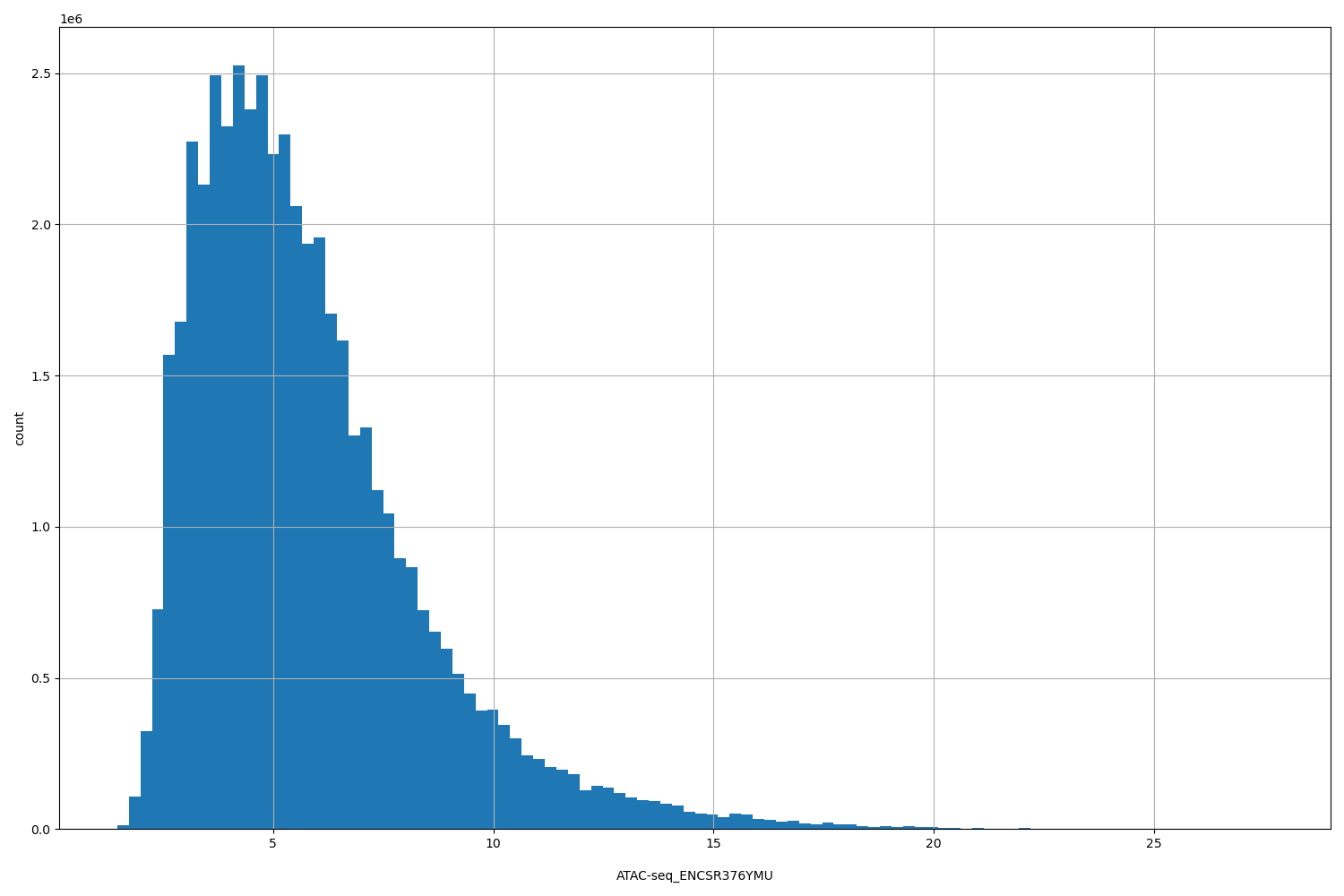
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR376YMU | float |
ATAC-seq_ENCSR376YMU |
ATAC-seq ENCSR376YMU [biosample_summary="Homo sapiens HG03439"]
|
 |
[1.46, 27.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF612NLR.bed.gz | 3.44 MB | c6162983c6e60969177cbd214caa5621 |
| ENCFF612NLR.bed.gz.dvc | 101.0 B | e9c2a466e54f670f60e66914b58fb5de |
| ENCFF612NLR.tabix.bed.gz | 981.25 KB | 127cdf3de507c51fd7e8eed1dd82ec03 |
| ENCFF612NLR.tabix.bed.gz.dvc | 107.0 B | 9340178a6b36cd73bde39c8d7d3e3082 |
| ENCFF612NLR.tabix.bed.gz.tbi | 422.65 KB | 6e1c9a635d4a3dfb96e63859cc12eaf0 |
| ENCFF612NLR.tabix.bed.gz.tbi.dvc | 110.0 B | e4221552f05bc0225fbcbca4673a0f52 |
| genomic_resource.yaml | 2.71 KB | 21e3eb63d508ecb7c3265f3a974cb96c |
| genomic_resource_original.yaml | 2.64 KB | 668ed1f0d005044f3601c9bf6deb1419 |
| statistics/ |