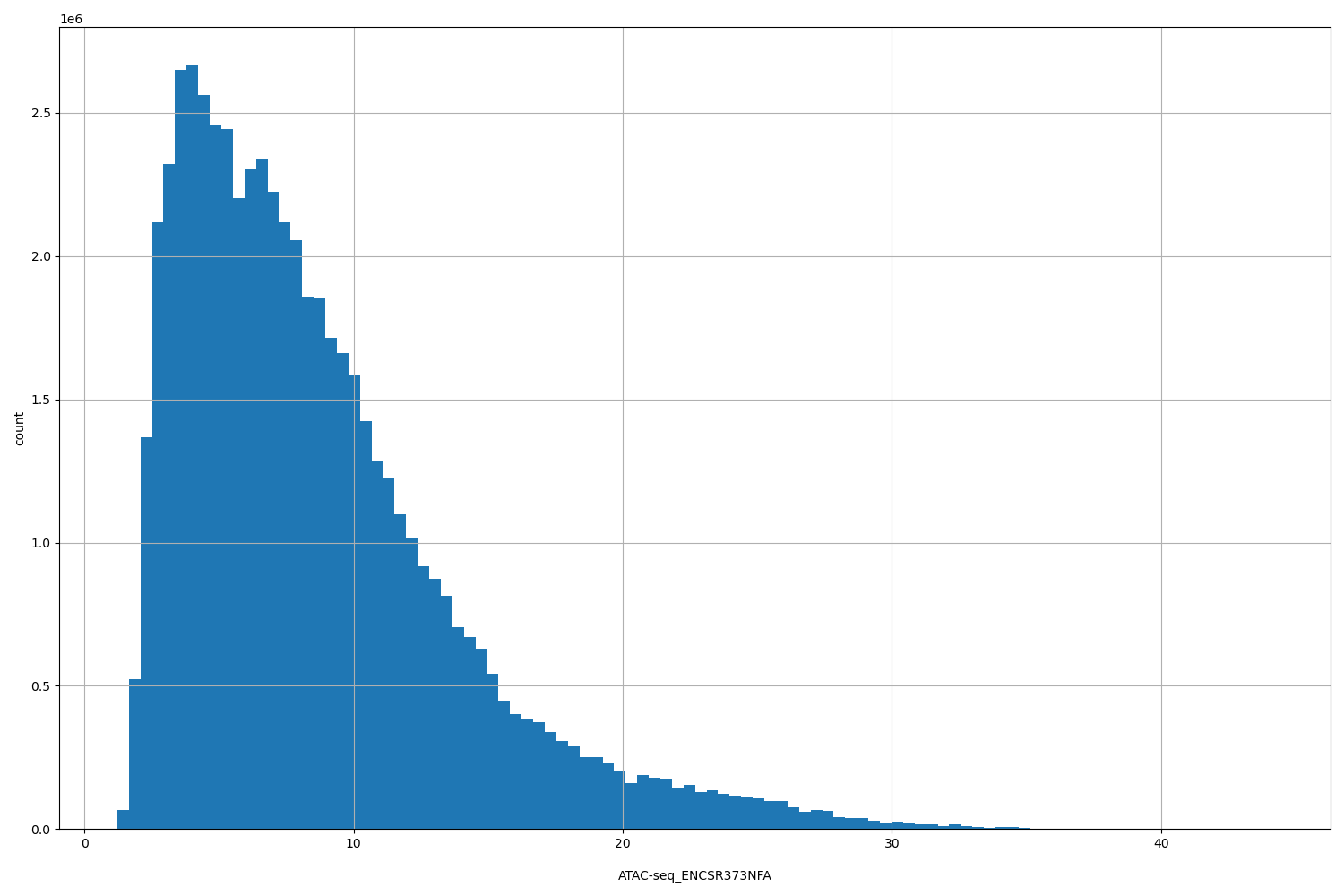
ATAC-seq_ENCSR373NFA
| Id: | ATAC-seq/ENCSR373NFA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR373NFA [biosample_summary="Homo sapiens activated naive CD4-positive, alpha-beta T cell male adult (50 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (50 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN455WBM|/analyses/ENCAN455WBM/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF913LRH|/files/ENCFF913LRH/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with NRF value of 0.877176. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF913LRH|/files/ENCFF913LRH/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with a PBC1 value of 0.89. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value < 0.7 is poor complexity, between 0.7 and 0.9 is moderate complexity, and >= 0.9 high complexity. NRF value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF151IFP|/files/ENCFF151IFP/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with NRF value of 0.876699. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.7 is acceptable. ENCODE processed file {ENCFF151IFP|/files/ENCFF151IFP/} processed by ATAC-seq ENCODE4 v1.10.0 GRCh38 pipeline was generated from a library with a PBC1 value of 0.89. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR373NFA | float |
ATAC-seq_ENCSR373NFA |
ATAC-seq ENCSR373NFA [biosample_summary="Homo sapiens activated naive CD4-positive, alpha-beta T cell male adult (50 years) treated with anti-CD3 and anti-CD28 coated beads for 72 hours, 50 U/mL Interleukin-2 for 72 hours"]
|
 |
[1.22, 44.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF792YKT.bed.gz | 3.88 MB | 57220ea5afa9922221079633991358b7 |
| ENCFF792YKT.bed.gz.dvc | 101.0 B | d861dd4272ced1809301c7ba12db9bcf |
| ENCFF792YKT.tabix.bed.gz | 1.12 MB | c0ba0634be075cf983313857f4c63653 |
| ENCFF792YKT.tabix.bed.gz.dvc | 107.0 B | 9ce64d3a572307e2bd72fcaf9d9f54c9 |
| ENCFF792YKT.tabix.bed.gz.tbi | 389.4 KB | 37a635528b3003f03ed7fca9609faadd |
| ENCFF792YKT.tabix.bed.gz.tbi.dvc | 110.0 B | 24740df669da18cb8dcea7962175969b |
| genomic_resource.yaml | 4.31 KB | 441859320c439c867f2ea556bb68cab8 |
| genomic_resource_original.yaml | 4.1 KB | 8494780dab493fe4b94b851e54bf08af |
| statistics/ |