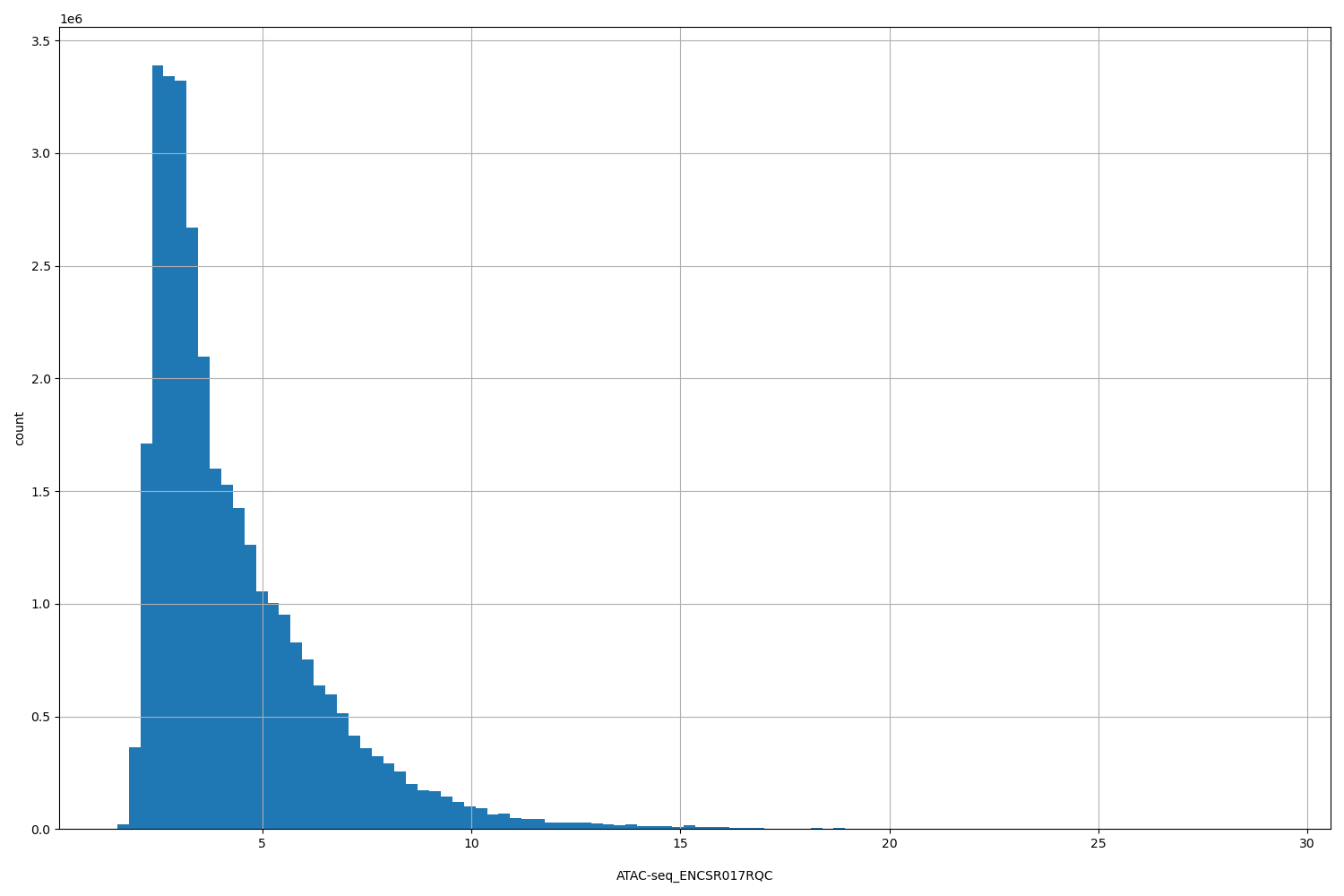
ATAC-seq_ENCSR017RQC
| Id: | ATAC-seq/ENCSR017RQC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
ATAC-seq ENCSR017RQC [biosample_summary="Homo sapiens Peyer''s patch tissue female adult (53 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (53 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN952JCC|/analyses/ENCAN952JCC/} has in progress subobject quality standard {encode4-atac-seq|/quality-standards/encode4-atac-seq/} audit_internal_action: Released analysis {ENCAN952JCC|/analyses/ENCAN952JCC/} has in progress subobject document {664cd382-ec9a-4478-994d-4f975424ad12|/documents/664cd382-ec9a-4478-994d-4f975424ad12/} audit_not_compliant: According to ENCODE4 standards, overlap peaks files in ATAC-seq assays processed by the uniform processing pipeline should have FRiP (fraction of reads in called peak regions) scores > 0.3. FRiP scores 0.2-0.3 are acceptable, and < 0.2 are not compliant. {ENCFF956UCP|/files/ENCFF956UCP/} processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline has a FRiP score of 0.08. audit_warning: According to ENCODE4 standards, ATAC-seq assays processed by the uniform processing pipeline should have either >150k reproducible peaks in an overlap peaks file, or >70k in an IDR thresholded peaks file. 100-150k or 50-70k peaks respectively is acceptable, and <100k or <50k respectively is not compliant. File(s) {ENCFF956UCP|/files/ENCFF956UCP/} (overlap peaks) and {ENCFF374BNC|/files/ENCFF374BNC/} (IDR thresholded peaks) processed by ATAC-seq ENCODE4 v1.9.1 GRCh38 pipeline have 101932 and 40947 peaks. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| ATAC-seq_ENCSR017RQC | float |
ATAC-seq_ENCSR017RQC |
ATAC-seq ENCSR017RQC [biosample_summary="Homo sapiens Peyer''s patch tissue female adult (53 years)"]
|
 |
[1.53, 29.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF956UCP.bed.gz | 2.32 MB | e025fac2b64f8f42b33d8912d3c95aa1 |
| ENCFF956UCP.bed.gz.dvc | 101.0 B | bb4dc20e04657704f05377ad5c27e5a7 |
| ENCFF956UCP.tabix.bed.gz | 750.03 KB | 43799fa9107d18736d01c0a2c4e186de |
| ENCFF956UCP.tabix.bed.gz.dvc | 106.0 B | 88cd8b7f6aedac94f24772a660412826 |
| ENCFF956UCP.tabix.bed.gz.tbi | 382.71 KB | f97ac5a77fe02102e078a54dfb9a9c06 |
| ENCFF956UCP.tabix.bed.gz.tbi.dvc | 110.0 B | a66b4bee73943cd4f7b8fcd0d631b8a3 |
| genomic_resource.yaml | 2.59 KB | c2a76ba00ba826639b7376f7445b3f5f |
| genomic_resource_original.yaml | 2.49 KB | fe09365b290cc93538dd18c2ac0b8373 |
| statistics/ |